Introduction
Peanut (Arachis hypogaea L.) is an important food legume and oilseed crop grown in many areas throughout the world. Diseases of peanut reduce yield and quality, and disease management efforts increase production costs wherever peanut is grown. In addition to increasing production costs, use of crop protective chemicals may result in environmental and food safety concerns. Therefore peanut breeding objectives must emphasize development of disease-resistant cultivars to help manage peanut diseases. In the USA, breeding for disease resistance was not given a high priority until the late 1970's because of the high value of the crop, the availability of chemicals for disease control, and a perceived lack of usable levels of resistance in cultivated germplasm (Wynne et al., 1991; Shew et al., 1995). Since then, numerous germplasm accessions among cultivated peanuts have been identified as sources of disease resistance (Holbrook and Dong, 2005; USDA, 2007). Additional resistant germplasm has also been identified among wild species collections (Stalker and Simpson, 1995; Holbrook, 2001).
Cylindrocladium black rot (CBR) caused by Cylindrocladium parasiticum Crous, Wingfield & Alfenas and Sclerotinia blight caused by Sclerotinia minor Jagger are economically important diseases in the Virginia-Carolina peanut production area. The diseases caused by these soilborne fungi are of great concern to growers and are difficult or expensive to control. CBR was first reported in North Carolina in the 1970's (Garren et al., 1972), where losses in individual infested fields can range from 1 to 50%. Statewide losses in Virginia and North Carolina are estimated at about 5% annually (Phipps, 2006). The first CBR-resistant cultivar, NC 8C, was released in 1983 (Wynne and Beute, 1983). Since the mid 1980's metam sodium has been used for soil fumigation (Phipps, 1990; Cline and Beute, 1986), providing partial control of the disease. In 2004, about 45% of NC peanut acreage was fumigated at a cost of approximately $75/ha (Brandenburg et al., 2005). CBR is seedborne (Randall-Schadel et al., 2001; Glenn and Phipps, 2003) at low rates (approx. 1.5%), so any existing population of inoculum in the soil can be augmented by new inoculum introduced on seeds. Therefore resistant cultivars are an important component of CBR management programs, along with crop rotation and soil fumigation. NC 12C (Isleib et al., 1997) and Perry (Isleib et al., 2003) are moderately resistant to CBR, but NC 12C is susceptible to Sclerotinia blight and Perry is susceptible to Tomato spotted wilt virus.
Approximately 25% of NC peanut fields in traditional production areas are infested with S. minor. Yield losses average about 5% per year, for a value at least $1 million annually (B.B. Shew, unpubl. data). Currently it is difficult to attain more than partial control with labeled fungicides (fluazinam and boscalid), which are very expensive (approximately $100 US/ha for each application). Early maturing cultivars VA 98R (Mozingo et al., 2000), Wilson (Mozingo et al., 2004), and NC-V 11 (Wynne et al., 1991b) tend to have less Sclerotinia blight damage due to earlier harvest and therefore less late season damage. The CBR-resistant cultivar, Perry, also has some resistance to Sclerotinia blight.
Developing cultivars with resistance to both diseases has been an objective of the breeding program at N.C. State Univ. However, field evaluations often fail to produce consistent results due to variation in weather within and across seasons and locations and the uneven distribution of soilborne inoculum (Hau et al., 1982; Porter et al., 1977). Therefore, greenhouse protocols were used to screen North Carolina breeding lines for resistance each winter from 2003 to 2006 and results were compared with performance of the lines in the field.
Materials and Methods
One hundred twenty-five breeding lines and check cultivars were tested in the field and greenhouse in at least one of the four years from 2003 through 2006; 51 genotypes were tested in at least two years; 34 were tested in at least three years, and 15 lines were tested in all four years. Of the genotypes tested, 107 were advanced breeding lines based solely on A. hypogaea germplasm, 8 were tetraploid (2n = 4x = 40) lines derived from interspecific hybridization of A. hypogaea with A. cardenasii Krap. & Greg., and 10 were released cultivars. Several lines derived from interspecific hybridization have been released as disease-, nematode- and insect-resistant germplasm (Stalker and Beute, 1993; Stalker et al., 2002a, b; Stalker and Lynch, 2002.) including one resistant to Sclerotinia blight (Isleib et al., 2006).
The Sclerotinia blight and CBR assays were performed in separate experiments under similar greenhouse conditions (23–27 C). All experiments were conducted as incomplete block designs with 4 replications for Sclerotinia blight tests and 6 replications for CBR tests.
Greenhouse Evaluation of Resistance to Sclerotinia Blight
Plants were grown in 10 cm pots containing a planting medium of two parts (v:v) steamed commercial topsoil to one part MetroMix 200 (Sun Gro Horticulture, Bellevue, WA) for 6 wk prior to inoculation. The inoculum was prepared by growing S. minor isolate P13, which was originally isolated from a diseased peanut in Chowan County, NC, on potato dextrose agar (PDA) for 2 days. A plug of PDA colonized by S. minor was placed in a 00 BEEM embedding capsule (Ted Pella, Inc., Redding, CA) with cap removed and was gently pushed onto a freshly cut petiole on the main stem of the plant. The first petiole from the bottom of the main stem that did not subtend a vegetative branch was used for inoculation. Inoculated plants were placed on a moisture-retaining mat on the greenhouse bench top and were misted 1 min every 2 hr during daylight hours but not at night. The bench area was enclosed on the sides and top with plastic sheeting over a PVC frame. After 48 hr the top cover was removed, but misting continued. Lesion length (mm) was measured 4, 5, 6, and 7 days after inoculation with a digital caliper (Empire Level MFG. Corp., Mukwonago, WI), and area under the disease progress curve (AUDPC) was calculated (Shaner and Finney, 1977).
Greenhouse Evaluation of Cbr Resistance
Two seeds of each genotypic entry were planted in a plastic cone-tainer, 3.81 cm dia and 20.96 cm in length (Stuewe & Sons, Inc, Corvallis, OR), with a cotton ball placed in the bottom to serve as a wick for water, then filled with a planting medium of two parts (v:v) steamed commercial topsoil, and one part MetroMix 200. The medium was artificially infested with 25 microsclerotia of C. parasiticum per g of medium at the time it was mixed. Four isolates of C. parasiticum originally obtained from diseased peanut grown in North Carolina were maintained in culture on PDA. Inoculum was prepared by transferring cultures to PDA and incubating at room temperature for 6 wk in the dark. Microsclerotia recovered from cultures of the four isolates were mixed in a water suspension and quantified (Black and Beute, 1984; Phipps et al., 1976).
Cone-tainers in racks were placed in plastic trays, and plants were grown for 8 wk in the greenhouse. Water in each tray was maintained at a height of approximately 10 cm (approx. 3 cm above the bottom of the tubes) to provide adequate soil moisture. The root system of any plant that died before harvest was washed and plated on PDA to determine whether C. parasiticum was present in the decaying roots. At harvest, surviving plants were removed from the cone-tainers, and the roots were washed and rated for degree of decay on a 0–5 scale (0 = no lesions or decay, 1 = few lesions on secondary roots and/or a few small lesions on taproot, 3 = many lesions on secondary roots and many lesions on the taproot and with several secondary roots missing, 5 = completely decayed roots with most secondary roots and part of taproot missing, with 2 and 4 = intermediate levels of severity (Black and Beute, 1984; Rowe and Beute, 1975). Any plant that died prior to the end of the experiment and was confirmed to have been infected with C. parasiticum received a score of 5. Any plant that died early, but did not harbor the fungus was considered a missing value.
Field Evaluations of Resistance
Field trials were conducted in farmers' fields in northeastern North Carolina from 2003 through 2006. Not all of the 125 genotypes were field-tested every year, but all genotypes were tested in at least one year. Each location was selected because it was reportedly infested with S. minor or C. parasiticum. At each location, there were three or more trials with some genotypes, particularly cultivars and resistant or susceptible checks, entered in more than one trial. In Sclerotinia blight trials, there was no application of preventive fungicides. In CBR trials, there was no fumigation with metam sodium. Leafspots and insects were controlled in all trials by application of fungicides and pesticides (Jordan et al, 2007). All trials were conducted as incomplete block designs with 3 or 4 replications. Fourteen seeds were planted in single-row plots 3.7 m in length with 25-cm seed spacing. Stand counts were made 4 to 6 wk after early May planting. Plants expressing symptoms of Sclerotinia blight were counted in mid- to late season dependent upon weather conditions favorable for disease, usually in late August or early September. Plants expressing symptoms of CBR were counted in late September or early October just prior to the time of normal harvest. Counts of diseased plants were converted to incidence scores, i.e., the proportion of symptomatic plants out of the number that emerged. Disease evaluation plots were not harvested.
Field Evaluations of Yield
From 2000 through 2006, field trials were conducted on three research stations [Peanut Belt Res. Sta. (PBRS) at Lewiston, Upper Coastal Plain Res. Sta. (UCPRS) at Rocky Mount, and Border Belt Tobacco Res. Sta. (BBTRS) at Whiteville] operated by the N.C. Dept. of Agriculture and Consumer Services. Several trials were conducted at PBRS and UCPRS each year but only one trial at BBTRS each year. Some genotypes appeared in more than one trial at a given location in a given year. Not all genotypes were tested at every location in every year. Sclerotinia blight was managed with labeled fungicides on demand. PBRS is the only one of the three stations known to harbor C. parasiticum, and all yield trials at that station were fumigated with metam sodium prior to planting. All other management practices were made according to the recommendations of the N.C. Coop. Ext. Ser. (Jordan et al, 2007). All experiments were conducted with two replications using square or rectangular lattice designs. Plots consisted of two rows 7.3 m in length and spaced 91 cm apart with a seed spacing of 25 cm. Stand counts were taken at approximately 4 wk after planting. Plots were dug at 145 to 155 days after planting, harvested by combine when sufficiently dried in the field, and further dried under forced heated air before measurement of yield. No disease data were recorded for these plots.
Methods Used for Data Analysis
Lesion lengths, AUDPC values, and root rot scores from the greenhouse experiments were subjected to analysis of variance for the incomplete block design employed in each year using the general linear models procedure (PROC GLM) of SAS v8.0 statistical software (SAS Institute, Cary, NC). If the effects of blocks within reps were not significant at P < 0.05, then blocks were dropped from the model and the data were analyzed as if they came from a randomized complete block experiment. Adjusted means were computed using the final model and included in the database for greenhouse experiments. The unbalanced set of adjusted means was subjected to analysis of variance in which the effects of years and lines were removed as sources of variation. Two analyses were performed: one of all 125 genotypes tested in any of the four years, and the second of the 15 experimental lines tested in all four years augmented by the six check cultivars tested in at least two out of the four years. Means were adjusted to a common year effect. All analyses were performed using PROC GLM of SAS statistical software.
Each year, data from the replicated field trials of disease reactions were analyzed and the adjusted means entered into a database from which subsequent summary analyses were performed. The unbalanced set of adjusted means from the database were subjected to analysis of variance in which effects of year, location, and lines were removed as sources of variation and means were adjusted to common environmental effects. A separate database of adjusted means for yield and market grade data was formed from trials conducted using recommended control measures for Sclerotinia blight and CBR since 2000 at PBRS, UCPRS and BBTRS. There was replication of genotypes from multiple tests from the three locations over time. Adjusted means from the unbalanced database were subjected to analysis of variance, and means adjusted to a common environmental effect were computed. All statistical analyses were performed using (PROC GLM) of SAS statistical software.
Correlation analyses were conducted to compare multiple-year means of greenhouse assays with disease incidence computed from field data. Pod yields of lines were also compared and correlations with disease data conducted.
Results
Petioles of plants inoculated with S. minor in the greenhouse exhibited water-soaked symptoms after 24 hr and lesions were found on stems by 3 to 4 days after inoculation. The inoculation method consistently produced lesions on more than 95% of the plants inoculated. In some cases, the petiole of an inoculated plant became water soaked, but no lesion developed. Occasionally, petioles and capsules fell off due to the weakening of petioles after attack by the fungus. Usually the fungus had already invaded the stem and stem lesions developed as usual. Otherwise, data for the plant was recorded as missing.
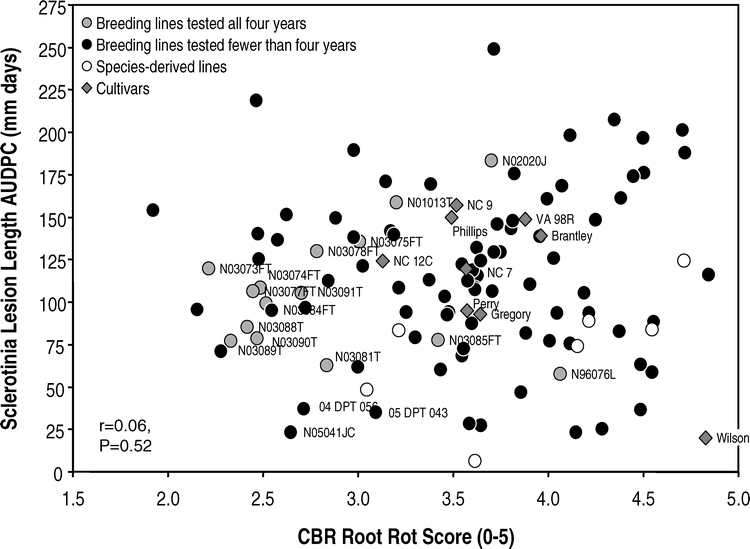
Of the 125 genotypes tested in the greenhouse in at least one of the four years, AUDPC for Sclerotinia blight ranged from 6.4 to 248.7 mm days for breeding lines and 20.0 to 157.0 mm days for cultivars (Fig. 1). For CBR root rot scores, the range was 1.9 to 4.8 for all lines and 3.1 to 4.8 for cultivars. Among the cultivars, Wilson had the lowest AUDPC for Sclerotinia blight lesion length while NC 12C had the lowest CBR root decay score. No cultivar exhibited low mean values for both disease reactions. There was no significant correlation between the two disease reactions (r = 0.06, P = 0.5237). Rank correlation was also not significant (r = 0.06, P = 0.4996).
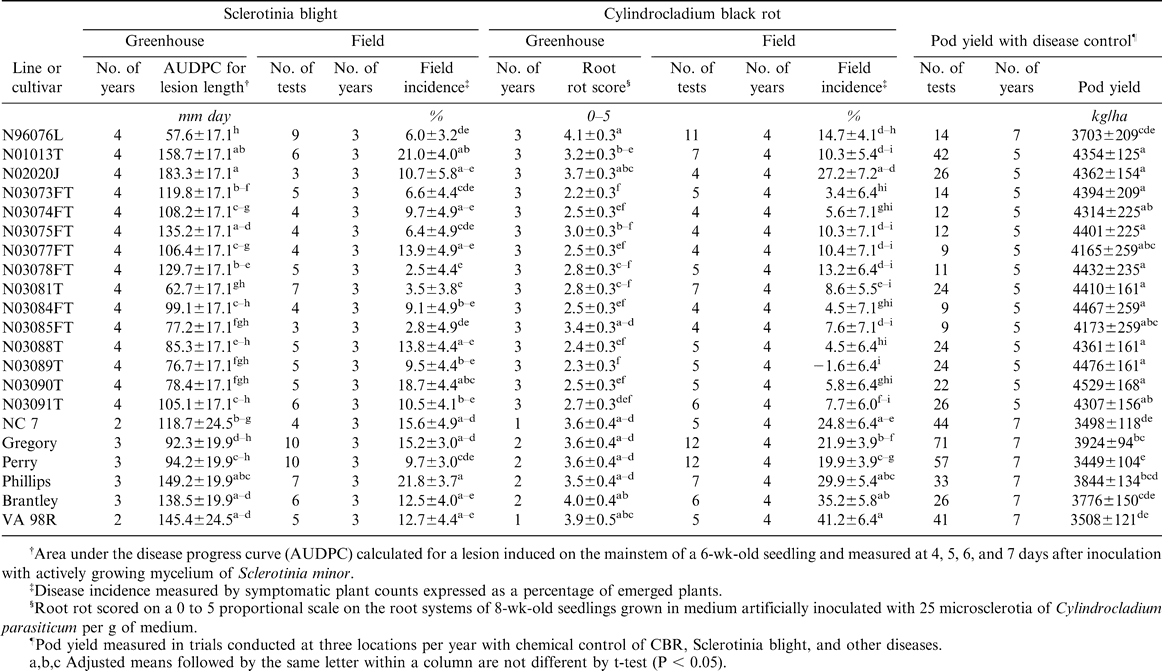
For the 15 experimental lines tested in the greenhouse assays in all four years, lesion length AUDPC ranged from 57.6 mm days for the germplasm line N96076L to 183.3 mm days for N02020J (Table 1). The corresponding range for check cultivars was 92.3 to 149.2 mm days. The contrast between experimental lines and checks was not significant (105.6 vs 123.1 mm days, P = 0.0843) unless N02020J, a line selected for large seed size rather than disease resistance, was grouped with the checks. The breeding line N03081T performed nearly as well as N96076L, with an AUDPC of 62.7 mm days.
Sclerotinia blight incidence in the field ranged from 2.5% on N03078L to 21.0% on N01013T (Table 1). Greenhouse and field results for Sclerotinia blight were not significantly correlated (r = 0.35, P = 0.1170). Several lines, for example N03090T and N03078FT, did not perform consistently in greenhouse inoculation vs. field trials. However, the most resistant lines in the greenhouse, N96076L and N03081T, also had low incidence of Sclerotinia blight in the field.
Adjusted means for CBR incidence in the field across four years ranged from −1.6% in N03089T to 27.2% in N02020J (Table 1). Greenhouse ratings of CBR resistance were highly correlated with CBR incidence in the field for the 15 lines tested in four years (r = 0.83, P < 0.0001). In addition to N03089T, several other lines performed well in the field and in the greenhouse trials. N96076L appeared somewhat less susceptible to CBR in the field in comparison to the greenhouse, where it was the most susceptible line tested.
N03090T had the highest yield at 4529 kg/ha (Table 1). All of the breeding lines tested produced higher yields than the germplasm line N96076L, which averaged 3703 kg/ha over the period of the study. The contrast between experimental lines and the checks was highly significant (4320 vs. 3766 kg/ha, P < 0.0001).
Discussion
The correlation of the Sclerotinia blight greenhouse assays with field incidence was 0.35 (P = 0.1170, n = 15) over all four years and 0.66 (P = 0.0080, n = 15) when the 2006 data was removed from the analysis. Incidence of Sclerotinia blight was low in 2006 resulting in a poor test to evaluate resistance that may have biased the correlation downwards. A previous study, comparing the results of a detached-leaf Sclerotinia blight assay to field incidence of Sclerotinia blight reported a similar nonsignificant correlation of r = 0.57 (P = 0.1070, n = 9) (Hollowell et al., 2003). Based on these results and the previous report, it appears that greenhouse screens for Sclerotinia blight resistance are not a substitute for field screening. Greenhouse screens measure only physiological resistance (Chappell et al., 1995), and it is possible that subtle differences among some lines may not be detected in the greenhouse assays of Sclerotinia blight resistance. However, good performance in the greenhouse assay can be used to supplement field results in the selection of lines for Sclerotinia blight resistance.
A comparison of multiple-year means from the greenhouse assays with disease incidence means computed from field data revealed a correlation of r = 0.83 (P < 0.0001) for CBR, suggesting that the greenhouse assay is a reasonably good predictor of field performance. Similar greenhouse screening methods were used in the early identification of CBR resistant germplasm in peanut (Pataky et al., 1983).
Considering the set of 125 genotypes tested in one or more years, the absence of negative correlation between reactions to Sclerotinia blight and CBR suggests that it should be possible to identify lines with resistance to both diseases. Several of the sister lines tested in all four years had low values for both. Several newer lines that were tested in fewer than four years had superior reactions, e.g., N05041JC, 04 DPT 056 and 05 DPT 043 (Table 1, Fig.1). Several lines that exhibited susceptible reactions to either Sclerotinia blight or CBR were dropped from the greenhouse testing program and also from consideration for release as cultivars.
The germplasm line N96076L has been released as a source of multiple disease resistance, including resistance to Sclerotinia blight (Isleib et al., 2006). However, N96076L is highly susceptible to CBR and does not have yield and quality comparable to commercial cultivars. In comparison to N96076L, the advanced lines N03081T, N03088T, N03089T, and N03090T scored an average of 1.5 CBR rating units lower, had similar levels of Sclerotinia blight resistance, and had superior yields. These lines also have superior agronomic traits and have potential for commercial release as cultivars with multiple disease resistance (Isleib, unpubl. data).
Acknowledgements
The authors thank the N.C. Peanut Growers Assoc., the N.C. Crop Improvement Assoc., the N.C. Foundation Seed Producers, Inc., the National Peanut Board, and the Everett Byrd Foundation for financial support for this research.
Literature Cited
Black M. C. and Beute M. K. 1984 Relationships among inoculum density, microsclerotia size, and inoculum efficiency of Cylindrocladium crotalariae causing root rot on peanuts. Phytopathology 74 : 1128 – 1132 .
Brandenburg R. L. , Jordan D. L. , Shew B. B. , Wilcut J. W. , and Toth S. J. 2005 Crop Profile for Peanuts in North Carolina Raleigh N.C. Coop. Ext. Serv., N.C. State Univ 33 pp. Revised. (http://www.ipmcenters.org/cropprofiles/docs/NCpeanuts.html).
Chappell G. F. , Shew B. B. , Ferguson J. M. , and Beute M. K. 1995 Mechanisms of resistance to Sclerotinia minor in selected peanut genotypes. Crop Sci 35 : 692 – 696 .
Cline W. O. and Beute M. K. 1986 Effect of metam sodium, peanut genotype and inoculum density on incidence of Cylindrocladium black rot. Peanut Sci 13 : 41 – 45 .
Garren K. H. , Beute M. K. , and Porter D. M. 1972 The Cylindrocladium black rot of peanut in Virginia and North Carolina. J. Am. Peanut Res. Educ. Assoc 4 : 66 – 71 .
Glenn D. L. and Phipps P. M. 2003 Incidence and survival of Cylindrocladium parasiticum in peanut seed. Plant Dis 87 : 867 – 871 .
Hau F. C. , Campbell C. L. , and Beute M. K. 1982 Inoculum distributions and sampling methods for Cylindrocladium crotalariae in a peanut field. Plant Dis 66 : 568 – 571 .
Holbrook C. C. 2001 Status of the Arachis germplasm collection in the United States. Peanut Sci 28 : 84 – 89 .
Holbrook C. C. and Dong W. 2005 Development and evaluation for a mini core collection for the U.S. peanut germplasm collection. Crop Sci 45 : 1540 – 1544 .
Hollowell J. E. , Shew B. B. , and Isleib T. G. 2003 Evaluating isolate aggressiveness and host resistance from peanut leaflet inoculations with Sclerotinia minor. Plant Dis 87 : 402 – 406 .
Isleib T. G. , Rice P. W. , Bailey J. E. , Mozingo R. W. , and Pattee H. E. 1997 Registration of ‘NC 12C’ peanut. Crop Sci 37 : 1976 .
Isleib T. G. , Rice P. W. , Mozingo R. W. , Bailey J. E. , Mozingo R. W. , and Pattee H. E. 2003 Registration of ‘Perry’ peanut. Crop Sci 43 : 739 – 740 .
Isleib T. G. , Rice P. W. , Mozingo R. W. , Copeland S. C. , Graeber J. B. , Shew B. B. , Smith D. L. , Melouk H. A. , and Stalker H. T. 2006 Registration of N96076L peanut germplasm. Crop Sci 46 : 2329 – 2330 .
Jordan D. L. , Spears J. F. , Brandenburg R. L. , Brown A. B. , Shew B. , Roberson G. T. , and Bullen S. G. 2007 2007 Peanut Information. N.C. Coop. Ext., Raleigh AG-331.
Mozingo R. W. , Coffelt T. A. , and Isleib T. G. 2000 Registration of ‘VA 98R’ peanut. Crop Sci 40 : 1202 – 1203 .
Mozingo R. W. , Coffelt T. A. , Swann C. W. , and Phipps P. M. 2004 Registration of ‘Wilson’ peanut. Crop Sci 44 : 1017 – 1018 .
Pataky J. K. , Black M. C. , Beute M. K. , and Wynne J. C. 1983 Comparative analysis of Cylindrocladium black rot resistance in peanut: greenhouse, microplot, and field testing procedures. Phytopathology 73 : 1615 – 1620 .
Phipps P. M. , Beute M. K. , and Barker K. R. 1976 An elutriation method for quantitative isolation of Cylindrocladium crotalariae microsclerotia from peanut field soil. Phytopathology 66 : 1255 – 1259 .
Phipps P. M. 1990 Control of Cylindrocladium black rot of peanut with soil fumigants having methyl isothiocyanate as the active ingredient. Plant Dis 74 : 438 – 441 .
Phipps P. M. 2006 Applied research on field crop disease control. Va. Polytechnic Inst. and State Univ. Info. Ser. No. 482. 152 pp.
Porter D. M. , Powell N. L. , and Cobb P. R. 1977 Severity of Sclerotinia blight of peanuts as detected by infrared aerial photography. Peanut Sci 4 : 75 – 77 .
Randall-Schadel B. L. , Bailey J. E. , and Beute M. K. 2001 Seed transmission of Cylindrocladium parasiticum in peanut. Plant Dis 85 : 362 – 370 .
Rowe R. C. and Beute M. K. 1975 Variability in virulence of Cylindrocladium crotalariae isolates on peanut. Phytopathology 65 : 422 – 425 .
Shaner G. and Finney R. E. 1977 The effect of nitrogen fertilization on the expression of slow-mildewing resistance in Knox wheat. Phytopathology 67 : 1051 – 1056 .
Shew B. B. , Beute M. K. , and Stalker H. T. 1995 Toward sustainable peanut production: progress in breeding for resistance to foliar and soilborne pathogens of peanut. Plant Dis 79 : 1259 – 1261 .
Stalker H. T. and Beute M. K. 1993 Registration of four leafspot-resistant peanut germplasm lines. Crop Sci 33 : 1117 .
Stalker H. T. and Simpson C. E. 1995 Germplasm resources in Arachis. 14 – 53 in: Pattee H. E. and Stalker H. T. Advances in Peanut Science Stillwater, OK Amer. Peanut Res. and Educ. Soc., Inc .
Stalker H. T. , Beute M. K. , Shew B. B. , and Barker K. R. 2002 Registration of two root-knot nematode-resistant peanut germplasm lines. Crop Sci 42 : 312 – 313 .
Stalker H. T. and Lynch R. E. 2002 Registration of four insect-resistant peanut germplasm lines. Crop Sci 42 : 313 – 314 .
Stalker H. T. , Beute M. K. , Shew B. B. , and Isleib T. G. 2002 Registration of five leaf spot-resistant peanut germplasm lines. Crop Sci 42 : 314 – 316 .
USDA, ARS, National Genetic Resources Program Germplasm Resources Information Network - (GRIN) [Online Database] Beltsville, Maryland National Germplasm Resources Laboratory URL: http://www.ars-grin.gov/cgi-bin/npgs/html/index.pl (23 January 2007).
Wynne J. C. and Beute M. K. 1983 Registration of NC 8C peanut. (Reg. No. 27). Crop Sci 23 : 184 .
Wynne J. C. , Beute M. K. , and Nigam S. N. 1991 Breeding for disease resistance in peanut (Arachis hypogaea L.). Annu. Rev. Phytopathology 29 : 279 – 303 .
Wynne J. C. , Coffelt T. A. , Mozingo R. W. , and Anderson W. F. 1991 Registration of ‘NC-V 11’ peanut. Crop Sci 31 : 484 – 485 .
Notes
- Agric. Res. Spec., Res. Asst. Prof. and Ext. Spec., Dept. of Plant Pathology, North Carolina State University, Raleigh, NC 27695-7903 [^]
- Prof., Sr. Researcher, Res. Asst., Dept. of Crop Sci., North Carolina State University, Raleigh, NC 27695-7629 [^] * Corresponding author - J.E. Hollowell
Author Affiliations