Introduction
Spotted wilt of peanut (Arachis hypogaea L.), caused by the tomato spotted wilt virus (TSWV), is currently one of the major factors limiting peanut yield in the U.S. In the Virginia-Carolina growing region, spotted wilt has gradually increased in severity since the mid 1990s. Incidence and damage of the disease in peanuts was the highest in both states during 2002 (Hurt et al., 2003).
Symptoms of spotted wilt in peanut are variable and include moderate to severe stunting, appearance of elaborate concentric ring spots on individual leaflets, bud and leaf necrosis, and even plant death (Ghanekar et al., 1979; Halliwell and Philley, 1974). The first symptoms of the virus usually appear a few weeks after planting, and newly symptomatic plants emerge thereafter for the remainder of the growing season. The growth stage at which the plant is infected determines the degree of yield reduction (Culbreath et al., 1992). Plants infected early in the season are the most affected, showing severe stunting and producing very few or no seed. However, reductions in both quantity and quality of pods and seed are also observed in plants infected at later growth stages (Culbreath et al., 1992).
TSWV is vectored in nature by several species of thrips (Thysanoptera) (German et al., 1992). The virus is acquired by immature thrips feeding on infected host plants and subsequent transmission takes place primarily through feeding activities of adults. TSWV has the ability to replicate within the vector, allowing it to transmit the virus for long periods of time. Therefore, viruliferous adult thrips are capable of infecting many plants (Ullman et al., 1993). Even though TSWV is vectored only by thrips, control of thrips with insecticide applications has proved ineffective in reducing the incidence of spotted wilt (Todd et al., 1994). There are few effective cultural and chemical practices for management of the disease (Culbreath et al., 1994). Although several factors have been shown to provide some suppression of disease, no single measure by itself has been effective under heavy disease pressure. Among all known factors that can be manipulated to reduce the risk of spotted wilt (including peanut cultivar, planting date, plant population, in-furrow insecticide application, and tillage practices), cultivar selection appears to have the most potential for minimizing the risk of losses to spotted wilt (Culbreath et al., 1999, 2000; Hurt et al., 2003). Although they vary in degree of susceptibility, none of the virginia-type cultivars released to date have a high level of field resistance, and they may suffer significant damage under intense epidemics. Cultivars with higher levels of resistance would be of great benefit across the Virginia-Carolina growing region. Moreover, cultivars are needed that combine TSWV resistance with good yield and quality.
Historically, plant breeders have been faced with the problem of selecting parents for the development of populations that have both high expected mean performance, and genetic variation for desirable traits. Identification of parental combinations that meet those two criteria increases the probability of recovering superior genotypes for cultivar development. The conventional method of selecting parents is based on their own performance. Observed performances are then used to calculate midparent values (MPV), or the mean of the parental means, to predict cross combination means. This method of parental selection poses some obvious disadvantages such as performance estimate biases when not all genotypes are evaluated or when data is missing in some environments (Panter and Allen, 1995b). Moreover, the efficiency of phenotypic selection in discriminating among superior individuals is reduced as the heritability decreases, and becomes very inefficient for traits with low heritability values (Falconer, 1989). Furthermore, performance testing of new genetic material is one of the most important and also most expensive aspects of plant breeding programs. Selecting superior lines is usually accomplished by testing a large group of lines across several locations and years. Statistical methods that maximize the accuracy of the estimate of performance of a line from fewer environments would be extremely useful for plant breeders (Panter and Allen, 1995b).
Henderson (1975) described the use of a mixed model to calculate the best linear unbiased predictors (BLUPs) of breeding values of potential parents based on observed data and the known variance-covariance structure among fixed and random effects. Genetic effects are considered to be random in the model while environmental effects are considered to be fixed. Henderson's method uses the degree of genetic relationship among individuals to determine the genetic structure of the population, and assumes that correlation between data on different individuals is caused only by additive genetic effects (Henderson, 1975). By using genetic relationships among individuals, related individuals contribute to the predicted values for one another. Information from relatives can contribute to the predicted breeding value for an individual for which there is little or no data. Moreover, the magnitude of that contribution is dependent on the degree of relationship between the two individuals (Panter and Allen, 1995a).
The BLUP procedure could be widely applicable in crop breeding programs because no additional experiments are required for obtaining the predictions. Instead, they are made from data that is routinely generated in a breeder's testing program, including performance data and estimates of genetic relationship among lines (Bernardo, 1996b). Best linear unbiased prediction has been widely used in livestock breeding (Henderson, 1975) and to a lesser degree, in forest tree breeding (White and Hodge, 1988). Among crop species, BLUPs have been used to estimate breeding values to identify superior cross combinations in maize (Bernardo, 1994, 1995, 1996a, b, c), soybean (Panter and Allen, 1995a, b), peach (de Souza et al., 1998a, b, 2000), sugarcane (Chang and Milligan, 1992), peanut (Pattee et al., 2001), and oil palm (Purba et al., 2001). The main goal of this study was to explore the use of the BLUP method for selection of lines with superior ability to transfer decreased TSWV incidence in combination with five other important agronomic and quality traits in peanut.
Materials and Methods
Experimental Materials
The material analyzed included 118 breeding lines from the N. C. State Univ. peanut breeding program, 12 virginia-type cultivars and one var. hirsuta (A. hypogaea subsp. hypogaea var. hirsuta Köhler) accession. Plants were grown and harvested using recommended procedures for peanut production in North Carolina. TSWV trials were conducted using wide plant spacing (25–51 cm between seeds) and no insecticide.
Evaluations
Spotted wilt was evaluated using a disease incidence rating where the number of severely stunted, chlorotic, wilted or dead plants was counted in each plot two times during the growing season. That number was then converted to a percentage of the total number of plants per plot. For TSWV incidence, genotypes were evaluated over 18 tests in 7 year-by-location combinations. Not all genotypes were included in all tests, so replication ranged from 1 to 15 tests with a mean of 3.
Data on yield (lb A−1), meat content (% of kernels from 500 g of clean unshelled pods), extra large kernels (% of extra large kernels, i.e. seeds that ride a 8.4 × 19.0 mm slotted screen, based on the 2002 federal grade sheet for virginia-type peanuts, from 500 g of clean unshelled pods), pod brightness (Hunter L scale), and crop value ($ A−1) were compiled. These data consisted of the lines' means from each test in which the line occurred. Because some lines had been tested for yield and quality more extensively than others, there was a wide range in the number of records for each line. In total, genotypes were evaluated for yield and quality over 84 tests in 30 year-by-location combinations.
Statistical Analysis
The mixed model procedure (PROC MIXED) in SAS (SAS Institute, 2001b) was used for the analysis of the unbalanced data set to calculate means for genotypes adjusted to a common environmental effect. The following additive genetic mixed model was used to predict the additive genetic effect for each individual:
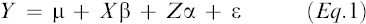 Eq.1
Eq.1
-
Y is a vector of observations,
-
β is a vector of fixed effects,
-
α is a vector of additive genetic effects,
-
ε is a vector of error terms, and
X and Z are incidence matrices that associate specific effects with individual observations.
The variance-covariance matrix for the random effects and error terms is
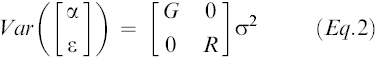 Eq.2
Eq.2
The standard BLUP solutions were obtained from the following equation
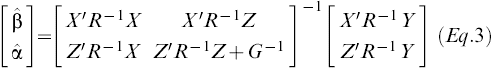 Eq.3
Eq.3
Heritability Estimates
Narrow-sense heritability (h2) estimates were not available for the overall breeding population for any of the six traits studied. However, it is known from quantitative genetics theory that the broad-sense heritability is the upper limit for the narrow-sense heritability. Estimates of broad-sense heritability (H) were calculated based on variance estimates obtained by restricted maximum likelihood estimation using PROC MIXED in SAS (SAS Institute, 2001b) and considering genotypic, year, location, and interaction effects to be random. BLUPs were calculated for a range of values around our estimates of H to assess the sensitivity of the method to inaccuracy in the estimation of narrow-sense heritability.
Selection Schemes
To select lines with superior breeding values for a combination of traits, independent culling and index selection were used as selection methods. For independent culling, a threshold value was chosen so that only the best 28-43% of the lines would be selected for a particular trait. For index selection, six different weighting schemes based on assigned importance of disease resistance vs. yield vs. agronomic and quality traits were designed. Subsequently, BLUPs were scaled as
 Eq.4
Eq.4
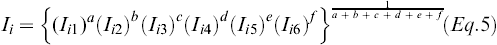 Eq.5
Eq.5
Results and Discussion
Heritability Estimates and their Effect on BLUP Values
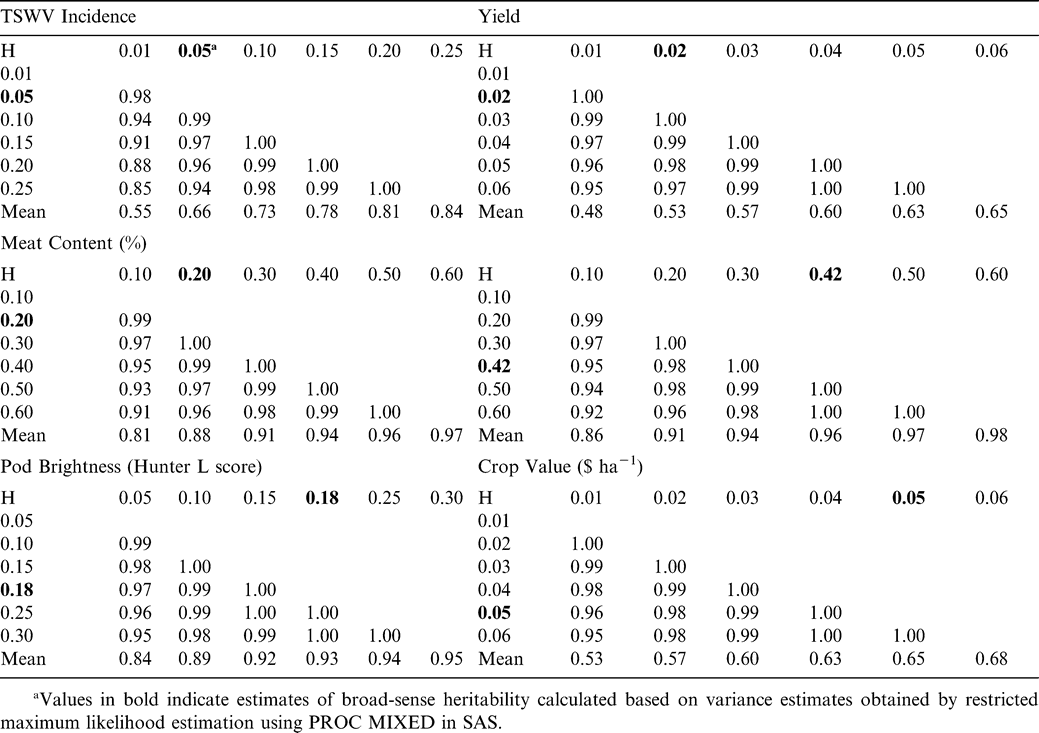
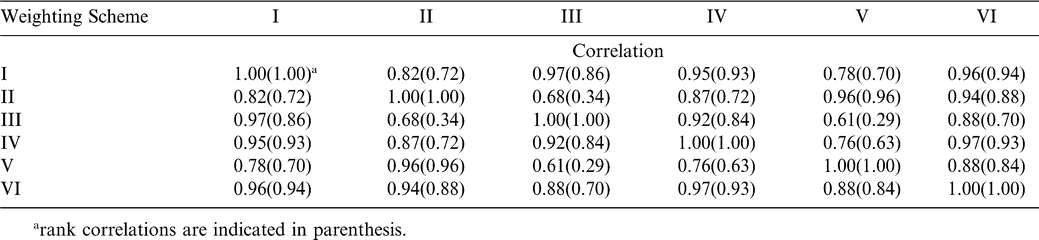
The additive variance-covariance matrix needed for BLUP estimation is based on estimates of narrow-sense heritability (h2). Only estimates of broad-sense heritability (H) were available for the six traits analyzed in this study. Only the additive variance is accounted for in h2, while H reflects all genotypic variance. Given that h2 must be less than or equal to H, BLUPs were calculated for a range of values around our estimates of H (Table 1). Subsequently, correlations among BLUPs calculated with different values of H were computed in order to investigate the sensitivity of the technique to variation in the heritability estimate. Correlations ranged from high to extremely high depending on the trait (r = 0.85 for TSWV incidence, r > 0.90 for all other traits). These results suggest that best linear unbiased prediction is relatively insensitive to inaccuracy in the estimation of narrow-sense heritability. Therefore, broad-sense heritability estimates can be used as substitutes without much loss in the estimation precision when estimates of narrow-sense heritability are not available (Pattee et al., 2001).
Correlation between BLUP Values and Means
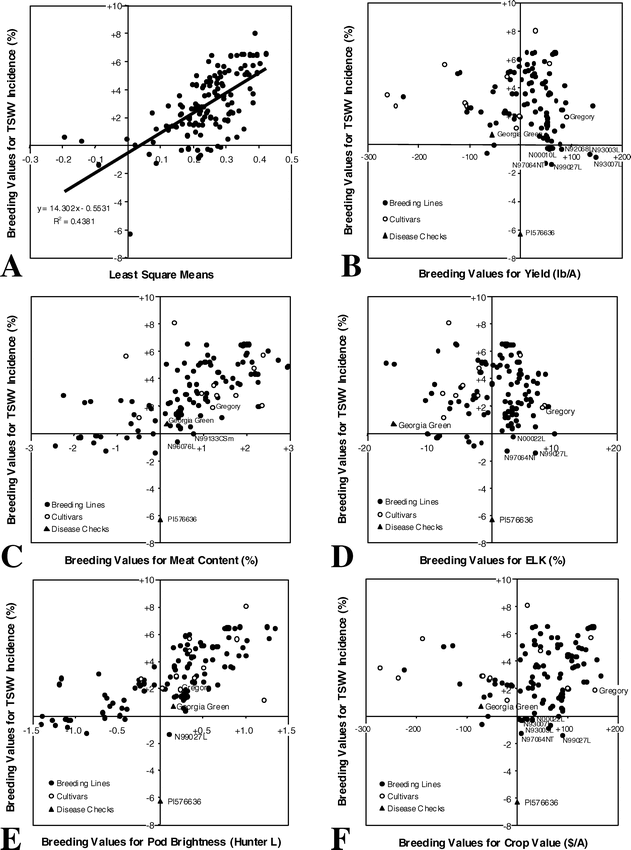
The use of phenotypic values to select parents should be effective in cases where the narrow-sense heritability is high (Falconer, 1989). However, for traits with low narrow-sense heritability values, breeding values (BV) would give a better ranking of the genetic value of the parents than would their phenotypic values, and, therefore, selection efficiency would be enhanced (de Souza et al., 2000). In this study, meat content, extra large kernels (ELK), and pod brightness had moderate broad-sense heritabilities of 0.20, 0.42, and 0.18, respectively. The predicted BVs of these three traits were well correlated (0.88, 0.96, and 0.93, respectively) with observed phenotypic values (Table 1). On the other hand, TSWV incidence, yield, and crop value, had very low broad-sense heritabilities (0.05, 0.02, and 0.05, respectively) and showed poor correlations between predicted BVs and observed phenotypic values (0.66, 0.53, and 0.65, respectively). For TSWV incidence, the plot of predicted BVs vs. means supports the low correlation between these parameters (Fig. 1A). Therefore, TSWV seems to be an ideally suited trait for parental selection based on BLUP estimation of BVs.
Best linear unbiased predictors (BLUPs) of breeding value for tomato spotted wilt virus (TSWV) incidence vs.: (A) least square means for TSWV incidence, (B) BLUPs of breeding value for yield, (C) BLUPs of breeding value for meat content, (D) BLUPs of breeding value for extra large kernels (ELK), (E) BLUPs of breeding value for pod brightness, and (F) BLUPs of breeding value for crop value, in Virginia-type peanuts.
Variation of BLUP Values
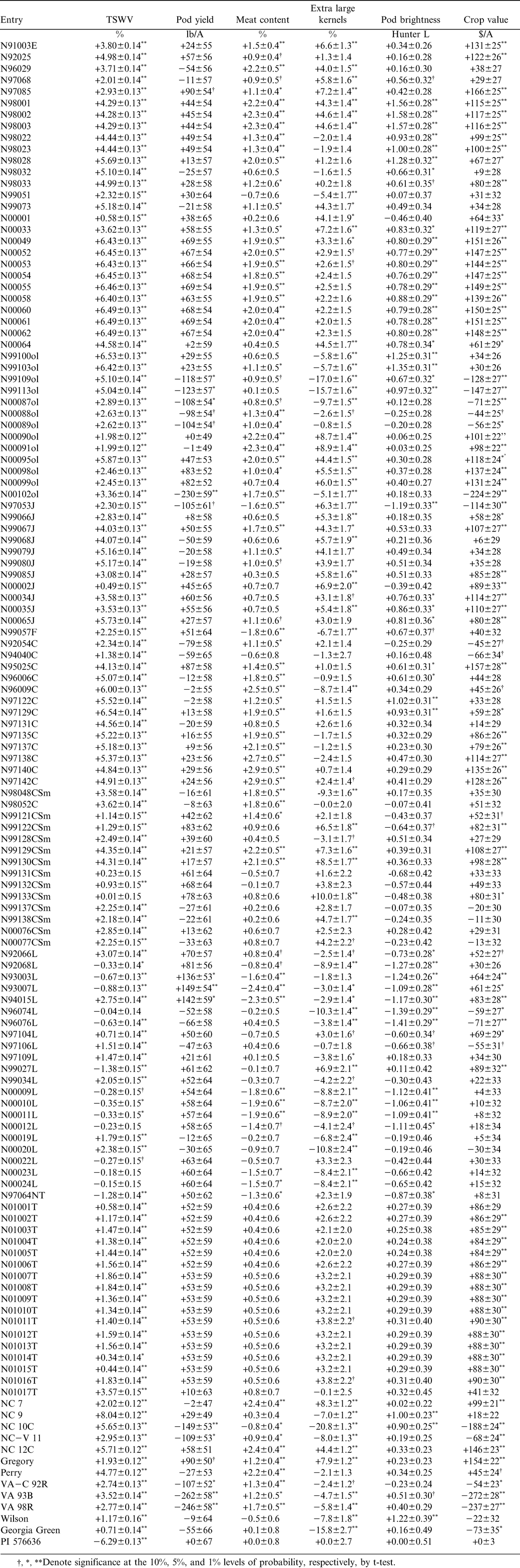
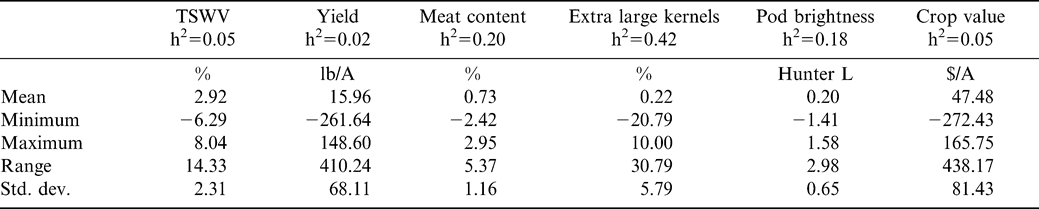
Best linear unbiased prediction was used to predict BVs of parents for TSWV incidence, yield, meat content, ELK, pod brightness, and crop value (Table 2). The predicted BV ranged from –6.29 to +8.04% for TSWV incidence, −262 to +142 lb A−1 for yield, −2.4 to +3.0% for meat content, −20.8 to +10.0% for ELK, −1.4 to +1.6 Hunter L units for pod brightness, and –272 to +166 $ A−1 for crop value (Table 3). Predicted BVs suggest that not only is there genetic potential to develop lines with increased field resistance to spotted wilt, but also that agronomic and quality traits can be improved.
Based on the BLUPs, several lines had superior (negative) BVs for TSWV incidence. A group of lines from our leafspot resistance breeding program had negative BVs for TSWV incidence, indicating that progenies from these lines would have reduced incidence of spotted wilt. Of the cultivars included in this study, none had negative BVs for TSWV incidence. Georgia Green and Wilson had the lowest positive values. Hirsuta accession PI 576636, a genotype with excellent field resistance to TSWV that was used as a resistant check in all tests, had the lowest BV for TSWV incidence among all genotypes analyzed. However, BVs for this accession might not be accurate due to its complete lack of genetic relationship to any other line in the data set.
For each agronomic and quality trait, several lines possessed an extremely high BV. However, no single line had the best BV for all the traits combined. Predicted BVs for TSWV incidence were plotted against those for yield to select lines that would combine negative BVs for TSWV incidence and positive BV for yield (Fig. 1B). A set of lines with resistance to leafspot and TSWV was found to possess superior BVs for both traits. An important point to highlight is how the BVs for cultivar Gregory for these two traits compare to those of other cultivars. Although its BV for TSWV incidence is slightly inferior to that of resistant cultivar Georgia Green, its BV for yield is considerably higher than that of any other cultivar.
Plots of predicted BVs for meat content and pod brightness against those for TSWV incidence indicate that only two lines, N96076L and N99133CSm, combine desirable BVs for meat content and TSWV incidence (Fig. 1C); and only one line, N99027L, combines desirable BVs for pod brightness and TSWV incidence (Fig. 1E). Several lines combined negative BVs for TSWV incidence and positive BVs for ELK and crop value (Figs. 1D and 1F).
Independent Culling
To select lines that would combine superior BVs for all traits analyzed, threshold values were selected that would pick the top 28−43% percent of the lines (top 28% for TSWV, top 41% for yield, top 43% for meat content, top 38% for ELK, top 35% for pod brightness, and top 28% for crop value). Subsequently, lines that had been picked for TSWV incidence, yield, and at least one of the four other traits were selected. Ten lines were selected including three (N99122CSm, N99132CSm, and N99133CSm) belonging to the Sclerotinia blight-CBR resistance breeding program, two (N99027L, and N00022L) to the early leafspot resistance breeding program, and five (N01009T, N01010T, N01011T, N01014T, and N01015T) to the TSWV resistance breeding program. These lines should transmit to their progenies not only reduced TSWV incidence, but also increased yields and improved quality traits.
Index selection
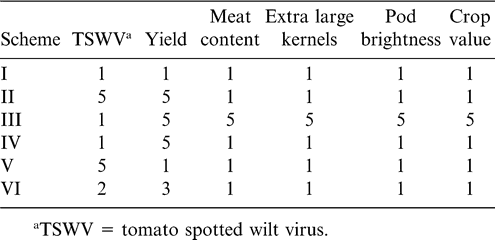
Weights of the different index selection schemes were based on assigned importance of disease resistance vs. yield vs. agronomic and quality traits (Table 4). The first scheme considered all traits to be equally important. Schemes II and VI emphasized TSWV and yield. Schemes III and V were reciprocal: III emphasized agronomic and quality traits, while V emphasized disease resistance. Scheme IV gave more importance to yield than to any of the other traits. Lines were ranked based on their index values. Subsequently, lines that had been ranked among the top 18 with at least four of the six weighting schemes were selected. Index values obtained using different weighting schemes were highly correlated (0.78 < r < 0.97) with the exception of schemes II and III (r = 0.68) and V and III (r = 0.61) (Table 6). Rank correlations were also found to be high (Table 5).
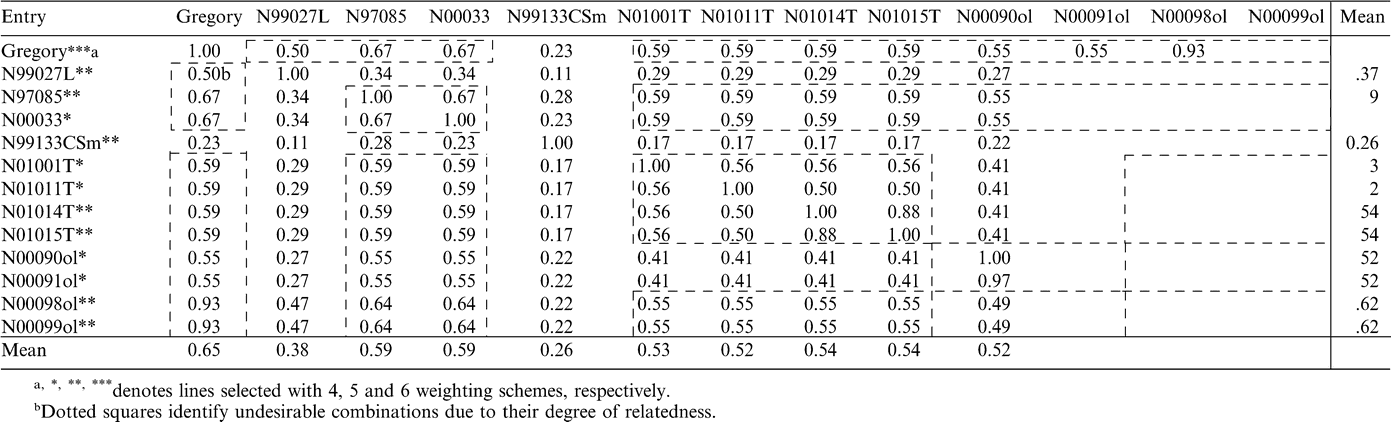
Thirteen lines were selected under each of four weighting schemes: N97085, N00033, N00090ol, N00091ol, N00098ol, N00099ol, N99133CSm, N99027L, N01001T, N01011T, N01014T, N01015T and Gregory. Of these, eight were selected under five weighting schemes and only one under all six schemes (Table 6). High oleic lines N00090ol, N00091ol, N00098ol, and N00099ol had excellent BVs for meat content and ELK. Moreover, N00098ol and N00099ol also had extremely high BVs for crop value. Although their BVs for TSWV incidence were positive, they were not large (Table 2). Likewise, TSWV lines N01001T, N01011T, N01014T, and N01015T had moderate positive BVs for TSWV incidence and good BVs for crop value. Leafspot line N99027L and CBR line N99133CSm had highly desirable BVs for TSWV incidence, but their BVs for agronomic and quality traits were not very high. Therefore, it would be valuable to utilize lines from the first set in crosses with lines from the second one to develop progenies that combine superior values for all traits. In doing so it is important to consider the degree of relationship between the lines to be crossed in order to have enough genetic variability in hybrid populations to allow additional improvement. Coefficients of coancestry among selected lines were examined in order to assess the amount of variability present among these genotypes (Table 6). Although some of the selected lines are closely related, enough variability remains within the group so as to continue genetic progress.
Surprisingly, none of the cultivars studied was chosen among the top 18 genotypes with any of the weighting schemes used with the exception of Gregory, which was the only genotype from the 131 analyzed to be selected with all six schemes. These results indicate that Gregory would be an excellent choice as a parent for an array of traits. This cultivar has the ability to transfer to its progeny good TSWV incidence, superior yield, and good values for meat content, ELK, pod brightness and crop value.
Application in Breeding Programs
The BLUP approach used in this study is the only procedure proven effective to predict single-cross performance (Bernardo, 1996a, b, c). Perhaps the most attractive feature of BLUP estimation is that no special experiments are required to obtain the predictions. Instead, the predictions are obtained by using data that is routinely generated in a breeder's testing program. Moreover, as more lines are tested in disease and/or yield trials each year, the effectiveness of the predictions will increase due to the larger number of observations that went into their estimation (Bernardo, 1996a).
Literature Cited
Bernardo R. Prediction of maize single cross performance using RFLPs and information from related hybrids. Crop Sci 1994 34 : 20 – 25 .
Bernardo R. Genetic models for predicting maize single-cross performance in unbalanced yield trial data. Crop Sci 1995 35 : 141 – 147 .
Bernardo R. Best linear unbiased prediction of maize single-cross performance. Crop Sci 1996a 36 : 50 – 56 .
Bernardo R. Best linear unbiased prediction of maize single-cross performance given erroneous inbred relationships. Crop Sci 1996b 36 : 862 – 866 .
Bernardo R. Best linear unbiased prediction of the performance of crosses between untested maize inbreds. Crop Sci 1996c 36 : 872 – 876 .
Chang Y. S. and Milligan S. B. Estimating the potential of sugarcane families to produce elite genotypes using bivariate prediction methods. Theor. Appl. Genet 1992 84 : 633 – 639 .
Cockerham C. C. Covariances of relatives from self-fertilization [Inbreeding, genetic components]. Crop Sci 1983 23 : 1177 – 1180 .
Culbreath A. K. , Todd J. W. , and Demski J. W. Productivity of Florunner peanut infected with tomato spotted wilt virus. Peanut Sci 1992 19 : 11 – 14 .
Culbreath A. K. , Todd J. W. , Branch W. D. , Brown S. L. , Demski J. W. , and Beasly J. P. Effect of new peanut cultivar Georgia Browne on epidemics of spotted wilt. Plant Dis 1994 78 : 1185 – 1189 .
Culbreath A. K. , Todd J. W. , Gorbet D. W. , Brown S. L. , Baldwin J. A. , Pappu H. R. , Holbrook C. C. , and Shokes F. M. Response of early, medium, and late maturing peanut breeding lines to field epidemics of tomato spotted wilt. Peanut Sci 1999 26 : 100 – 106 .
Culbreath A. K. , Todd J. W. , Gorbet D. W. , Brown S. L. , Baldwin J. A. , Pappu H. R. , and Shokes F. M. Reaction of peanut cultivars to spotted wilt. Peanut Sci 2000 27 : 35 – 39 .
de Souza V. A. B. , Byrne D. H. , and Taylor J. F. Heritability, genetic and phenotypic correlations, and predicted selection response of quantitative traits in peach: I. An analysis of several reproductive traits. J. Amer. Soc. Hort. Sci 1998a 123 : 598 – 603 .
de Souza V. A. B. , Byrne D. H. , and Taylor J. F. Heritability, genetic and phenotypic correlations, and predicted selection response of quantitative traits in peach: II. An analysis of several fruit traits. J. Amer. Soc. Hort. Sci 1998b 123 : 604 – 611 .
de Souza V. A. B. , Byrne D. H. , and Taylor J. F. Predicted breeding values for nine plant and fruit characteristics of 28 peach genotypes. J. Amer. Soc. Hort. Sci 2000 125 : 460 – 465 .
Falconer D. S. Introduction to Quantitative Genetics 1989 London Longman Sci. and Technol 3rd ed.
German T. L. , Ullman D. E. , and Moyer J. M. Tospoviruses: Diagnosis, molecular biology, phylogeny, and vector relationships. Annu. Rev. Phytopathol 1992 30 : 315 – 348 .
Ghanekar A. M. , Reddy D. V. R. , Izuka N. , and Gibbons R. W. Bud necrosis of groundnut (Arachis hypogaea) in India caused by tomato spotted wilt virus. Annals of Applied Biology 1979 93 : 173 – 179 .
Halliwell R. S. and Philley G. Spotted wilt of peanut in Texas. Plant Dis. Reptr 1974 58 : 23 – 25 .
Henderson C. R. Best linear unbiased estimation and prediction under a selection model. Biometrics 1975 31 : 423 – 447 .
Hurt C. , Brandenburg R. , Jordan D. A. , Shew B. B. , Isleib T. G. , Linker M. , Herbert A. , Phipps P. , Swann C. , and Mozingo R. W. Managing tomato spotted wilt virus in peanuts in North Carolina and Virginia. North Carolina Coop. Ext. Serv. Bull 2003
Malécot G. Les Mathémathiques de l'Hérédité 1948 Paris Masson et Cie .
Panter D. M. and Allen F. L. Using best linear unbiased predictions to enhance breeding for yield in soybean: I. Choosing parents. Crop Sci 1995a 35 : 397 – 405 .
Panter D. M. and Allen F. L. Using best linear unbiased predictions to enhance breeding for yield in soybean: II. Selection of superior crosses from a limited number of yield trials. Crop Sci 1995b 35 : 405 – 410 .
Pattee H. E. , Isleib T. G. , Gorbet D. W. , Giesbrecht F. G. , and Cui Z. Parent selection in breeding for roasted peanut flavor quality. Peanut Sci 2001 28 : 51 – 58 .
Purba A. R. , Flori A. , Baudouin L. , and Hamon S. Prediction of oil palm (Elaeis guineensis Jacq.) agronomic performances using the best linear unbiased predictor (BLUP). Theor. Appl. Genet 2001 102 : 787 – 792 .
SAS Institute SAS/IML User's Guide, Version 8 2001a Cary, NC SAS Institute Inc .
SAS Institute SAS/STAT User's Guide, Version 8 volumes 1, 2, and 3 2001b Cary, NC SAS Institute Inc .
Todd J. W. , Culbreath A. K. , Rogers D. , and Demski J. W. Contraindications of insecticide use relative to vector control of spotted wilt disease in peanut. Proc. Amer. Peanut Res. Educ. Soc 1994 26 : 42 (abstr.) .
Ullman D. E. , German T. L. , Sherwood J. L. , Westcot D. M. , and Cantone F. A. Tospovirus replication in insect vector cells: Immunocytochemical evidence that the nonstructural protein encoded by the S RNA of tomato spotted wilt tospovirus is present in thrips vector cells. Phytopathology 1993 83 : 456 – 463 .
White T. L. and Hodge G. R. Best linear prediction of breeding values in a forest tree improvement program. Theor. Appl. Genet 1988 76 : 719 – 727 .
Notes
- Dept. of Crop Science, Box 7629, N.C. State University, Raleigh, NC 27695-7629 [^] *Corresponding author - S.R. Milla-Lewis