Introduction
Marker-assisted breeding is a powerful tool for crop improvement. A set of genetic markers can be applied to a segregating population to assist the selection of individuals possessing genomic regions controlling the expression of desirable phenotypic traits. Defining the marker-trait associations for most agronomically important traits is challenging since these traits often are inherited quantitatively and can be influenced by environment. Multiple genetic resources have been utilized to determine genetic marker-trait associations. Quantitative trait mapping using biparental recombinant inbred line (RIL) populations is most widely adopted (Bernardo, 2008). A RIL population can be maintained in perpetuity and allows for measurements of multiple phenotypic traits across locations and years. Coincident quantitative trait loci (QTL) identified across multiple environments are considered reliable for the trait of interest (Collard et al., 2005). The limitation of biparental RILs lies in the relative narrow genetic base represented by the two parents and the limited number of recombination events that occur during meiosis of F1 hybrids. The second type of genetic resource that has been utilized for QTL discovery is genome-wide association (GWA) mapping populations which takes advantage of historical genetic recombination events that occurred across many generations leading to divergence among landraces and germplasm. However, the rate of identifying false QTL using GWA is relatively high due to unknown genetic structure or relatedness of the genetic materials. The third type of genetic resource is multi-parental populations such as nested association mapping (NAM) populations and multi-founder advanced generation inter-cross (MAGIC) populations. These structured populations combine the advantages of both RIL and GWA by having multiple parents, which enlarges the genetic pool and increases recombination events leading to higher resolution of genetic maps (Ladejobi et al., 2016). However the cost of managing this type of population is escalated due to its large size. A NAM population is constructed by breeding a number of diverse inbred founder lines with one reference line and advancing all cross combinations as RILs (Guo et al., 2010). NAM populations were established in Arabidopsis thaliana (L.) Heynh. (Buckler and Gore, 2007) and maize (Zea mays L.) (McMullen et al., 2009). In the case of the NAM population in maize, thousands of recombination events were captured which facilitates the genetic mapping of plant morphological and disease resistance traits (Buckler et al., 2009; Kump et al., 2011; McMullen et al., 2009).
Cultivated peanut is an allotetraploid and has a relatively narrow genetic base due to the recent polyploidization event that gave rise to this species (Bertioli et al., 2016). Genetic mapping has been particularly challenging due to the scarcity of genetic markers. A number of biparental RILs segregating for response to disease and abiotic stress, maturity and crop quality traits were established for cultivated peanut (Pandey et al., 2012). Mapped traits from peanut RILs include disease resistance (Khera et al., 2016; Pandey et al., 2016; Pandey et al., 2017; Sujay et al.; 2012), plant morphology (Zhou et al., 2016) and response to abiotic stress (Faye et al., 2015; Varshney et al., 2009). Most of these published QTLs cover large genomic regions due to the limitations of marker density and map resolution due to small mapping population sizes. The lack of precision in mapping QTL regions reduces the effectiveness of marker-assisted breeding and hampers map-based cloning for genes controlling phenotypic traits. To increase the resolution of genetic mapping and refine QTL regions, a NAM population was created in peanut (Holbrook et al., 2013). Two common parents, Tifrunner (Holbrook and Culbreath, 2007) and Florida-07 (Gorbet and Tillman, 2009), were selected as females and crossed with eight unique male parents: N08082olJCT, C76-16, NC 3033 (Beute et al., 1976), GP-NC WS 16 (Tallury et al., 2014), SSD 6 (PI 576638), OLin (Simpson et al., 2003), New Mexico Valencia A (Hsi and Finkner, 1972), and Florunner (Norden et al., 1969) resulting in 16 structured populations. Selection of these parental lines was the fruition of the peanut research community's aim to maximize genetic diversity and target traits important to the US peanut breeding programs. Major traits associated with the selected parental lines were reported previously (Holbrook et al., 2013). To characterize the genetic potential of this NAM parental germplasm, the first objective of this study was to identify and quantify traits that are likely to segregate among the populations. To this end, 25 traits related to yield, pod maturity, seed germination, plant morphology, disease resistance and salt tolerance were quantified among the parental lines. Traits demonstrating significant differences among the population parents were identified. The second objective was to determine the genetic polymorphism among the population parents. SNPs between population parents ranged from 1,000 to 4,000 on Version 1 and 2,000 to 9,000 on Version 2 arrays.
Materials and Methods
Field trials were conducted from year 2012 to 2014 at the Gibbs Farm and Lang-Rigdon Farm, University of Georgia, Tifton Campus, Tifton, Georgia. Ten parental genotypes were planted following a randomized complete block design with three field replicates. Each plot consisted of two 1.5 m rows seeded at 20 seed per m. The width of the plot was 0.9 m. Plots were managed throughout the growing season using standard production practices (Beasley, 2013; UGA, 2018) and were irrigated as needed.
Plot yield and pod traits
Upon harvest, peanuts were dried to less than 10%; moisture level. Yield of field plots was determined in 2014 for all genotypes. Hundred pod weight, hull weight from 100 pods and 100 seed weight were measured in 2012. Ten mature double-seeded pods per field replication were selected based on endocarp color of pods for each genotype (Williams and Drexler, 1981). Mature double-seeded pod weight, seed weight, and pod area were measured in 2012 and 2014. Pod area was determined from pod images using the Image J software (Wu et al., 2015). The numbers of single-seeded, double-seeded and triple-seeded pods were counted in 250 grams of randomly selected pods from each line. Percentage of each type of pod was calculated in 2014.
Pod maturity
Pods harvested at 140 d after planting (DAP) were blasted with a pressure washer. Mesocarp color at the saddle area of a double or triple-seeded pod was compared to the peanut profile board (Baldwin and Beasley, 1990) to determine maturity. Percentage of pods in the category of black (14 d to maturity) was determined for each genotype. Data for pod maturity were collected in 2013, 2014 and 2015.
Disease ratings
TSWV disease rating at the Gibbs farm in Tift County, Georgia was performed by documenting spotted wilt disease incidence in each plot at 105 DAP in 2013; 112 DAP in 2014; 90 DAP in 2015. The rating scale was proportional with a score of 0 indicating no symptomatic plants, 1 indicating 10%; symptomatic plants; 2 indicating 20%; symptomatic plants etc. A score of 10 indicated 100%; symptomatic plants in the plot (Tillman et al., 2006). Late leaf spot disease visual ratings were taken at 100, 114, 125 and 139 DAP at the Rigdon farm, in Tift County, Georgia in 2014 using the Florida scale (Knauft et al., 1988). Area under the disease progress curve (AUDPC) was calculated for late leaf spot disease ratings.
Plant morphology
Measurements of plant height and primary lateral length were taken from three plants per field plot for each genotype. The number of nodes on primary laterals was counted and average internode length was calculated by lateral length/number of nodes. Data collection was performed at 120 DAP and 138 DAP in 2013; 120 and 140 DAP in 2014; and 140 DAP in 2015. No statistically significant differences were detected between the two measurement dates in both 2013 and 2014. Therefore average data of the two harvest dates was taken for each year.
Germination test
Forty-five seeds per genotype were pre-treated with fungicide 5,6-dihydro-2-methyl-1,4-oxathiin-3-carboxamide (Vitavax at 3 g/kg seed, Crompton, Middleburry, CT) and germinated in water saturated germination paper for 7 d at 28 C. Images of germinated seeds were scanned for radicle length and root length measurements using the Assess Software Version 2.0 (Lamari, 2008).
Salt tolerance test
A salt tolerance test was performed in 2016. Seeds were germinated and grown for 5 to 7 d in petri-dishes (Fisher Scientific, Hampton, NH) lined with moistened filter paper (Fisher Scientific, Hampton, NH). For each treatment, 10 seedlings/genotype were transferred to seed germination pouches (mega-international.com) containing 20 ml of Hoagland's medium (Atamian et al., 2012) and kept in a growth chamber maintained at 25 to 28 C and 16 h light/8 h dark cycle for 10 d. Peanut seedlings were transferred to new seed germination pouches containing 20 ml of 0 mM, 100 mM and 200 mM NaCl, dissolved in water, for 6 d before measurements of the following parameters were taken: tap root length, whole root length, shoot fresh weight and root fresh weight. Shoots and roots were collected and dried in an incubator at 80 C for 3 d before the dry weights of shoots and roots were measured. All values were normalized to the average of no salt treatment controls for each genotype.
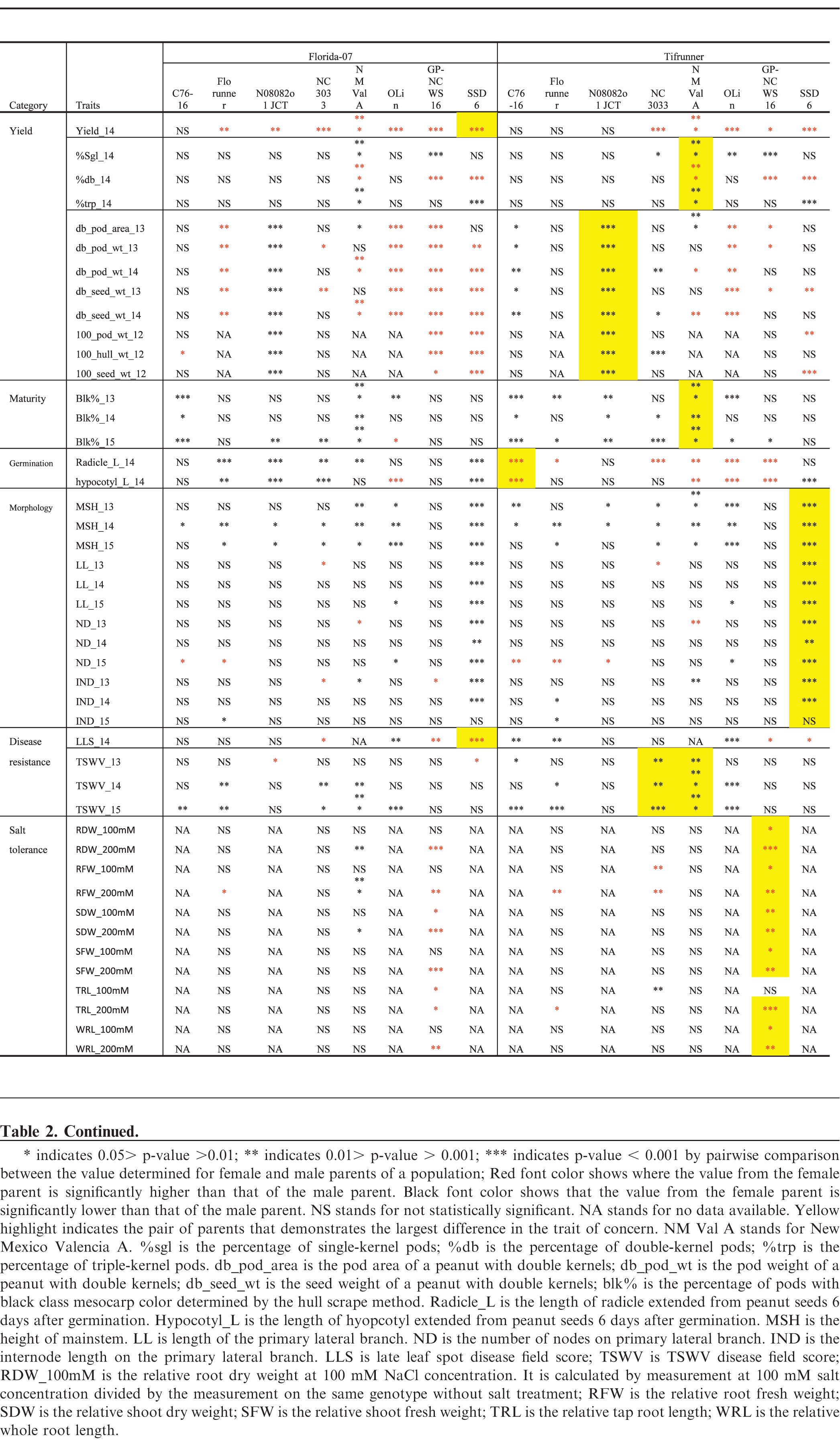
Phenotyping data were analyzed using mixed-model analysis of variance (PROC MIXED) of SAS using restricted maximum likelihood (REML) as the estimation model (SAS Version 9.4, SAS Institute). Effects of genotype, year, and genotype x year interaction are presented in Table S1. Significant genotype x year interaction showed differences in the performance of genotypes measured across the years. Therefore, when the genotype x year interaction was significant, the yearly data were analyzed separately. Significant differences between population parents were identified when the p-value was less than 0.05.
Genotyping
DNA was extracted from young leaves pooled from 10 to 15 individual plants per genotype using a DNeasy plant mini kit (Qiagen, Valencia, CA) and quantified by the Quant-it PicoGreen dsDNA assay kit (ThermoFisher Scientific, Pittsburgh, PA). Genotyping was performed on the Arachis Axiom SNP array Version 1 (Clevenger et al., 2017) and Version 2 by Affymetrix (ThermoFisher Scientific). SNP calling was performed with Axiom Analysis Suite (Version 1.1). Version 1 SNP number was curated manually to recover SNPs from all five categories 1) PolyHighResolution in which the samples were clearly grouped in three clusters and all of the samples passed quality control. 2) NoMinorHom in which all samples passed quality control yet only two clusters were observed. 3) OfftargetVariant in which additional low intensity clusters were present due to the slight mismatch between the probe and sample DNA. 4) CallRateBelowThreshold in which genotype call rate is below 90%;. 5) Other in which the clustering pattern does not conform to the previous categories. In the process of manual curation, the clustering profiles of all polymorphic markers were inspected. Markers demonstrating undesirable clustering profiles were excluded. Since the polymorphic marker density increased significantly with the Version 2 chip, only PolyHighResolution and NoMinorHom categories were output directly from the software.
Results and discussion
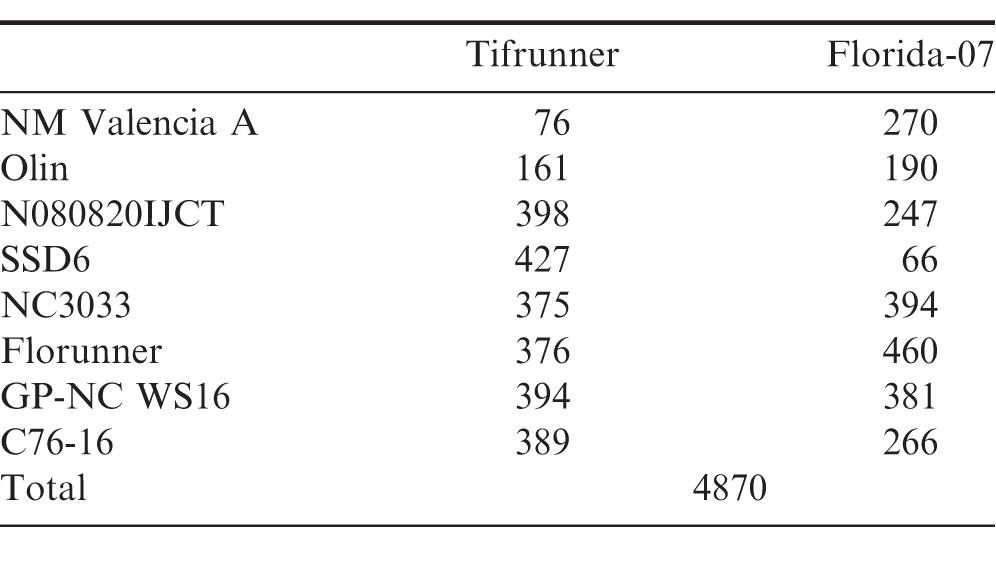
The total number of recombinant inbred lines for the NAM population is 4,780 (Table 1). The initial goal for each sub-population was 400, yet a few populations with New Mexico Valencia A, OLin and SSD6 as male parents had significantly fewer RILs due to attrition from disease pressure. In order to provide a guideline to prioritize phenotyping efforts for such a large population, we phenotyped the complete set of population parents.
Yield and yield components
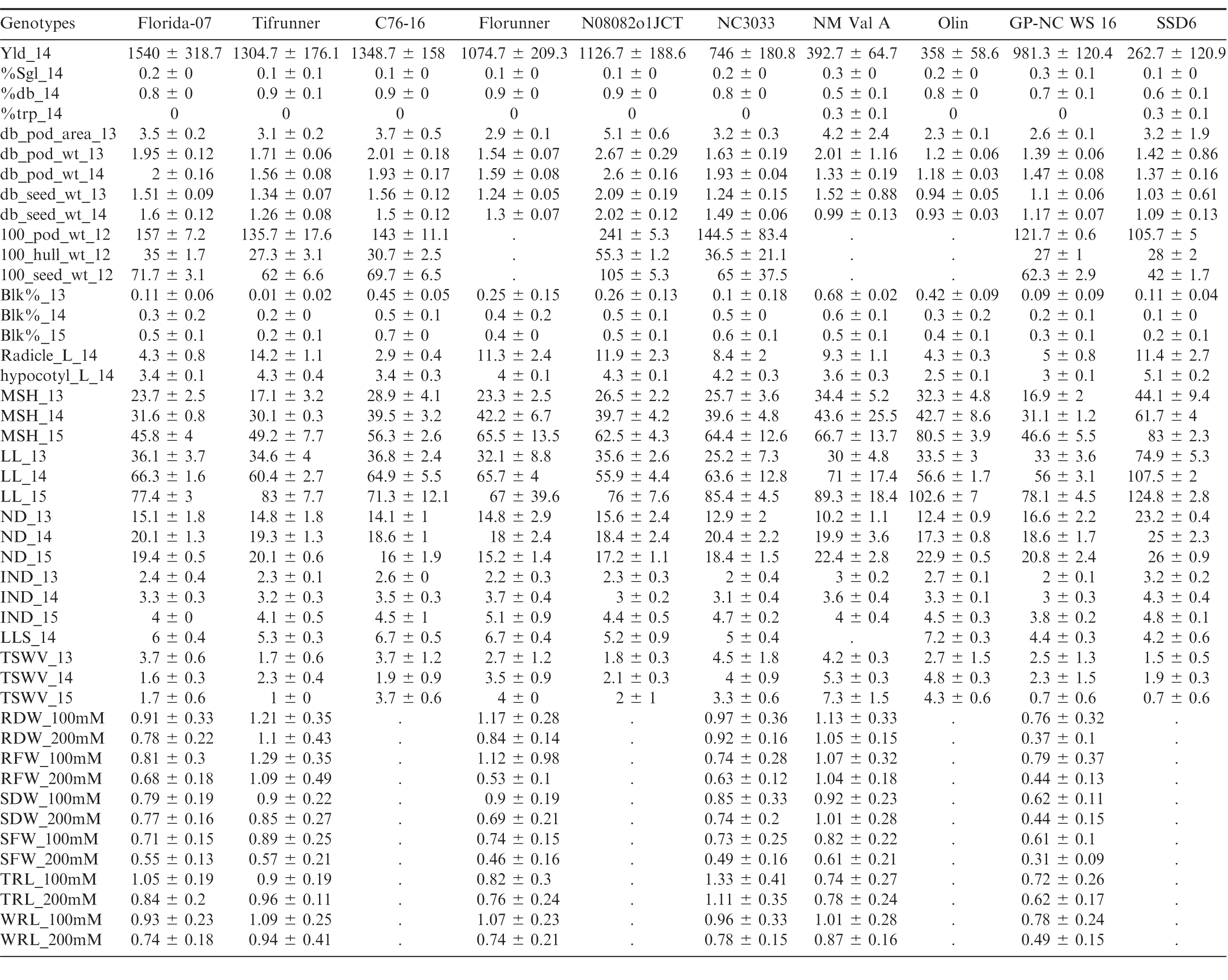
Florida-07 had significantly higher small-plot yield than all of the male parents except for C76-16 whereas the yield of Tifrunner was greater than NC3033, GP-NC WS 16, OLin, New Mexico Valencia A and SSD 6 (Tables 2, 3). The largest yield difference occurred between Florida-07 and SSD 6. Significant yield advantage of Florida-07 and Tifrunner is expected when growing these parental lines together in South Georgia. Although high yield is the ultimate goal for a breeding program, it is a quantitatively inherited trait influenced by multiple genetic factors in addition to the cumulative effects from farming practice, disease pressure and weather conditions. Therefore, genetic mapping of yield alone would be insufficient for marker development. Since number of seeds per pod, seed weight, and pod maturity are measurable yield components demonstrating heritability and interdependence in genetic mapping (Timmerman-Vaughan et al., 2005), these traits were evaluated among the population parents.
Number of seeds per pod
The number of seeds per pod is determined by the number of ovules initiated and at the seed morphogenesis stage when ovule abortion can occur in legumes (Ney et al., 1993). It is a yield component positively affecting peanut production. New Mexico Valencia A and GP-NC WS 16 had significantly higher percentages of single-kernel and fewer double-kernel pods than both female parents (Tables 2). SSD 6 and New Mexico Valencia A had significantly higher percentages of triple-kernel pods than either female parent. In order to map for the ratio of single, double and triple-kernel pods, both populations of Florida-07 x New Mexico Valencia A and Tifrunner x New Mexico Valencia A would be desirable since the parents demonstrated significant differences in all three measurements. New Mexico Valencia A originally was released for the production of three to four kernel pods (Hsi and Finkner, 1972). Previously, the number of seeds per pod was mapped in soybean to a 0.7 cM genomic region containing the Ln-encoding gene (Glyma20g25000.1) which was proposed to be the candidate gene (Jeong et al., 2011; Jeong et al., 2012). Mapping of seeds per pod in rapeseed also led to discovery of a major QTL explaining 32%; of phenotypic variation (Shi et al., 2015). Further genetic mapping of seeds per pod in peanut will shed light on the genetic control of this important agronomic trait.
Pod size and weight
Multiple measurements were taken to determine pod size and weight. Hundred-pod and hundred-seed weight are used by peanut breeders to determine the size of peanut; however, this measurement can be confounded by the maturity of pods because the pods are randomly sampled. It is known that pod maturity affects seed and pod size in peanut (Williams et al., 1987). Selection of mature double-kernel pods for weight and size measurement is a refined, although labor intensive, phenotyping procedure, intended to evaluate maximum potential. Male genotypes N08082o1JCT and OLin had the largest and smallest pods and seeds among all tested genotypes, respectively, in all environments (Tables 2, 3). Both Florida-07 and Tifrunner are significantly different from N08082o1JCT and OLin in pod and seed measurements. The largest difference among the population parents lies between Tifrunner and N08082o1JCT since N08082o1JCT is a large-seeded virginia-type peanut breeding line even larger in seed size than Bailey (Isleib et al., 2011). Florida-07 is a large-seeded runner type peanut and was significantly larger than GP-NC WS 16 in all measured parameters. Mapping the populations with N08082o1JCT and OLin as male parents will lead to discovery of additional QTLs conditioning pod and seed size traits in peanut.
Pod maturity date
Peanut is an indeterminate crop producing fruit on a single plant with a wide range of maturity levels. Pod maturity affects multiple aspects of peanut quality including flavor (Sanders et al., 1989), lipid profile (Sanders, 1980), and aflatoxin contamination (Dorner et al., 1989). Early pod maturity may help the crop to escape late season drought and diseases; however, late maturity was reported to be associated with higher tolerance to leaf spot disease (Branch and Culbreath, 1995; Miller et al., 1990). The two female parents are mid- to late-maturity cultivars which mature in 140 to 150 days after planting in South Georgia. The percentage of pods at full maturity, determined by the mesocarp color of pods reaching the black stage using the hull-scrape method, was measured at 140 days after planting in 2013, 2014 and 2015. C76-16 and New Mexico Valencia A demonstrated consistently earlier maturity than both Florida-07 and Tifrunner (Tables 2, 3). The greatest difference among population parental combinations was between Tifrunner and New Mexico Valencia A. Mapping this trait in the segregating subpopulations should help in pyramiding desirable genes for maturity with disease resistance.
Radicle and hypocotyl length of young seedlings
Roots and stems are vital parts of a plant for absorbing and transporting water and nutrients from soil. Root morphology and size are associated with plant tolerance to drought stress. Heritable root length differences among peanut varieties at an early growth stage were reported (Ketring et al., 1982). In this study, radicle and hypocotyl length were measured at 7 d post-germination, a time frame in which seedling stand counts may be affected. Active radicle and hypocotyl extension contributes to seedling vigor (Biswas et al., 2000). Tifrunner had significantly longer radicle and hypocotyl length than that of C76-16, New Mexico Valencia A, OLin and GP-NC WS16. The largest parental difference was found between Tifrunner and C76-16. C76-16 was reported to be a drought tolerant line due to its large root system when the drought stress was imposed late in the growing season (Dang et al., 2012). The radicle and hypocotyl length measured in this study only assessed root growth at the seedling stage, a different stage of root development than the previous report.
Plant morphology
Plant architecture is regulated genetically through hormonal signaling pathways (Sussex and Kerk, 2001). Main stem height has been used to describe the growth habit of cultivated peanut and is often documented in cultivar releases. Peanut forms vegetative and reproductive nodes on lateral branches following an alternating pattern in the subspecies hypogaea and sequential reproductive pattern in the subspecies fastigiata (Bunti ng, 1955; Smartt, 1961). Shorter internode length potentially could increase the productivity of peanut by resulting in more pods per unit length of the lateral branch. Excessive vegetative growth of peanut is considered unfavorable due the cost of management and photosynthate partitioning away from seed production (Bauman and Norden, 1971). Primary lateral length, node number on primary laterals and internode length data were collected in 2013, 2014 and 2015 (Tables 2, 3). SSD 6 had the longest main stem, primary laterals and highest number of nodes among all genotypes throughout three years of this study. Highly significant differences were found between SSD 6 and the two female parents, Florida-07 and Tifrunner. As for internode length, 2013 and 2014 demonstrated significant differences of this male parent from the two female parents. Data from 2015 did not reach statistical significance although the mean value of SSD 6 was higher. Populations with SSD 6 as male parent should be useful to map these morphological traits of peanut.
Late leafspot and TSWV resistance
Late leaf spot (caused by Cercosporidium personatum (Berk. & Curt.) Deighton) is a major fungal disease plaguing peanut production worldwide. It occurs mid-Aug in South Georgia, US and progresses until the end of the growing season, causing severe defoliation in susceptible genotypes. TSWV (genus Tospovirus, family Bunyaviridae) was first found in the southeastern US in the 1980s and soon became the main disease constraining peanut production (Culbreath and Srinivasan, 2011). Integrating host resistance and implementing effective field management are critical for yield protection. Among the parental genotypes, GP-NC WS 16 and SSD 6 are the best genetic resources for late leaf spot resistance (Tables 2, 3). Both genotypes have significantly lower disease scores than either Tifrunner or Florida-07. GP-NC WS 16 is a germplasm line with the leaf spot resistant diploid species, A. cardenasii Krapov. & W.C. Gregory, in its pedigree (Company et al., 1982). Resistance to late leaf spot was mapped by QTL-seq in the Florida-07 x GP-NC WS 16 population (unpublished data). Both Florida-07 and Tifrunner demonstrated resistance to TSWV compared to most males in this study (Tables 2, 3). TSWV field disease pressure varied across the tested environments and years. The most susceptible male parents were New Mexico Valencia A and NC3033.
Salt tolerance
Salinity stress from salt accumulation in soil reduces plant growth by increasing toxic metabolites, generating reactive oxidative species, and preventing photosynthesis and nutrient uptake (Jain et al., 2006). Salt tolerance is a desirable trait for peanut growing regions with high soil salinity levels. In order to test if there was any differential response to salt among the parental lines, young peanut seedlings were subjected to salt stress at 100 and 200 mM NaCl concentrations. GP-NC WS 16 had significantly reduced root and shoot growth compared to Tifrunner measured by root and shoot dry and fresh weights, whole root and tap root lengths at both salt concentrations (Tables 2, 3). Therefore, Tifrunner demonstrated better salt tolerance at the young seedling stage than GP-NC WS 16.
Genotyping data
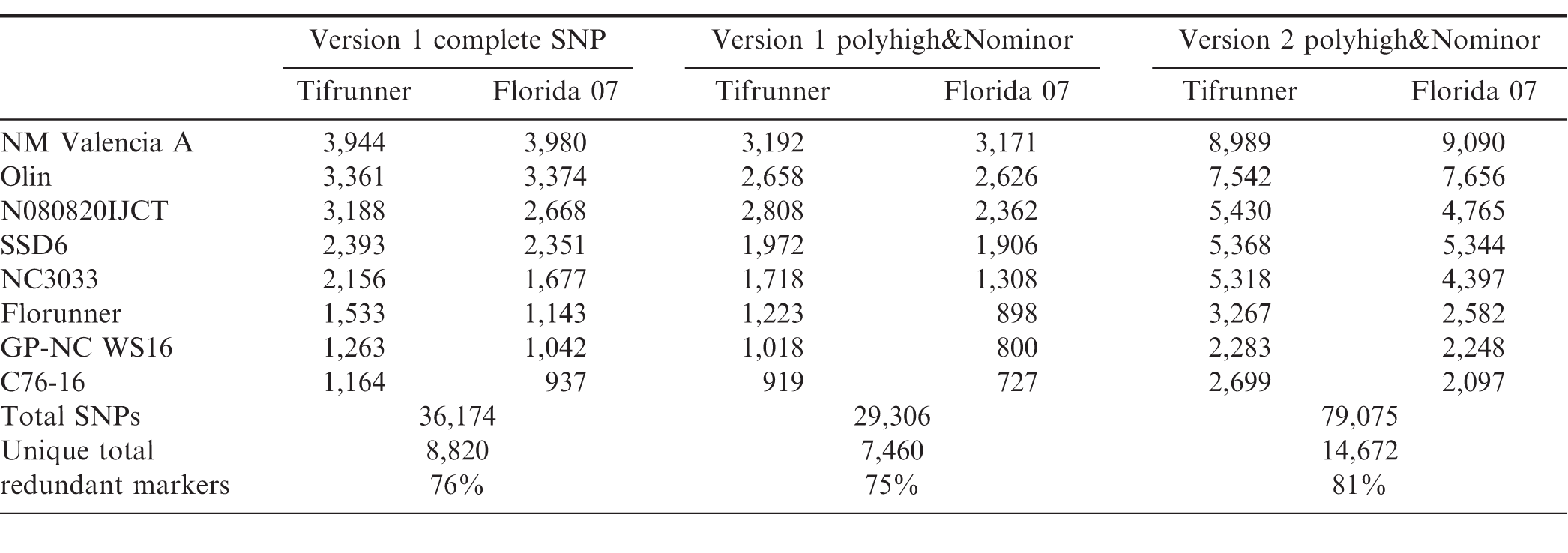
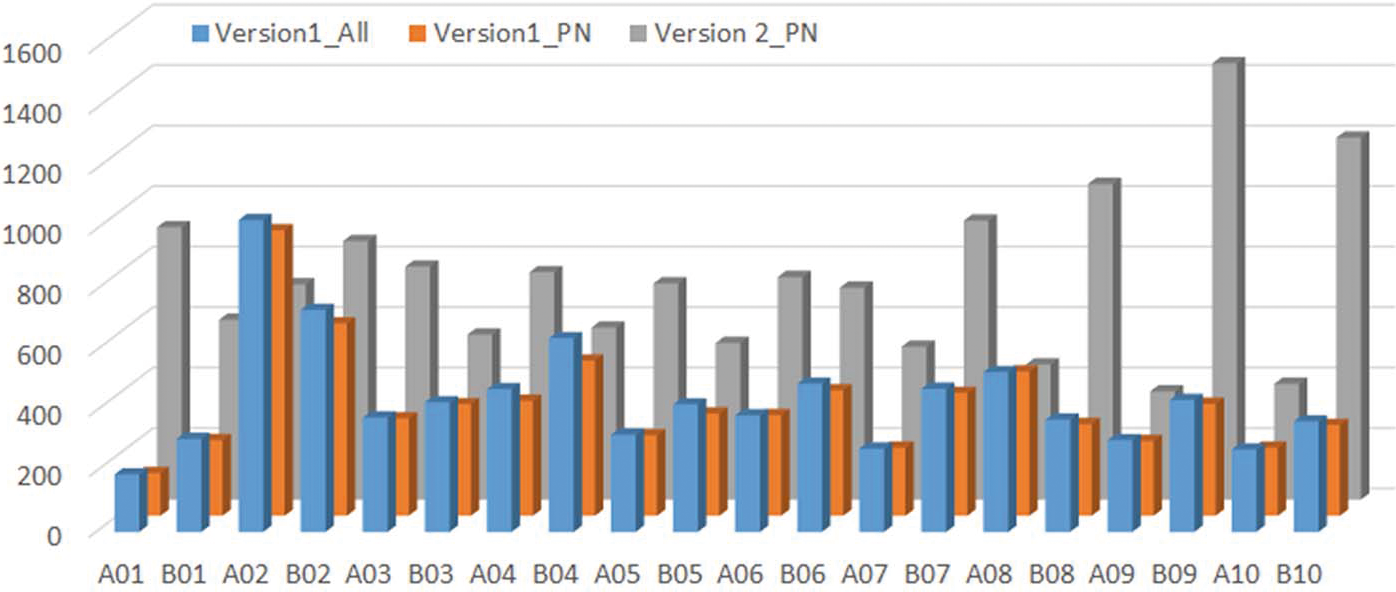
Genotyping platforms have evolved rapidly in recent years (Rasheed et al., 2017). Discovery of single nucleotide polymorphic (SNP) markers from next generation sequencing data greatly increased the marker density for peanut (Clevenger et al., 2015; Clevenger and Ozias-Akins, 2015). The construction of Affymetrix genotyping arrays with 58,233 probe sets on a Version 1 Arachis array (Clevenger et al., 2017) and 47,837 probe sets on a Version 2 array has enabled genotyping of mapping populations in a high throughput fashion. The Version 2 array integrated SNPs identified by an improved bioinformatics pipeline which will be described elsewhere. The number of Version 1 polymorphic markers between NAM population parents ranged from 937 to 3,980 with the lowest marker density found in Florida-07 x C76-16 and highest marker density in Florida-07 x New Mexico Valencia A (Table 4). The variation in range of marker density indicated level of genetic diversity among the parental lines. New Mexico Valencia A is genetically most distant from the runner-type peanut cultivar Florida-07 among all parental combinations. The total number of SNPs reached 36,174 for the sixteen biparental populations and there were 8,820 non-redundant SNPs across the NAM. Markers shared among populations were considered redundant. When only PolyhighResolution and NoMinorHom categories were taken into account on the Version 1 array, the total number of SNPs was reduced to 29,306 which accounted for 81%; of the total SNPs. The remaining 19%; of markers recovered from CallRateBelowThrshold, OffTargetVariant and Other categories were manually curated since the SNP data quality was too low for high confidence automated calling in these categories. The second version of the array has increased the number of subgenome specific SNP markers; therefore, the output of polymorphic markers from PolyhighResolution and NoMinorHom categories avoids manual calling and resulted in 79,075 total SNPs among the population parents. This number doubled the total number of SNPs from Version 1 and was 2.7 times the output of the same two categories from Version 1 data. Non-redundant SNPs among population parents was 14,672 from the Version 2 array. A majority (76 to 81%;) of the SNPs from Version 1 and Version 2 arrays were redundant which provides common markers to perform meta-analyses and build a consensus map for the 16 NAM sub-populations. The highest marker density was found on chromosome A02 and the lowest on chromosome A01 on the Version 1 array (Figure 1). Besides chromosomes A02 and A08, marker density on the remaining 18 chromosomes increased on the Version 2 array. The largest increase of marker density was found on chromosome A01 in which Version 2 had five times more markers than that of Version 1 followed by B10 and B09 (3 times more).
Number of SNP markers detected among NAM parents using the Arachis Axiom SNP array Version 1 and Version 2. All categories columns include SNPs from all categories including PolyhighResolution, NoMinorHom, OffTargetVarient, CallRateBelowThreshold and Other. Polyhigh&Nominor columns include SNP markers from polyhighresolution and Nominorhom categories only.
The distribution of SNP markers identified from version 1 and version 2 arrays across chromosomes. Version1All is the number of SNP markers from all categories of SNP markers. Version 1PN and Version 2PN are the number of SNP markers from polyhighresolution and Nominorhom categories on Version 1 and 2 chips only.
Conclusions
In summary, contrasting phenotypic data from plant morphology and disease resistance to pod and seeds traits were collected for the 10 parental lines of the NAM population, and up to 79,000 SNP markers were identified among the parental genotypes. Further genotyping of the NAM sub-populations using the Affymetrix SNP arrays will allow the construction of high density genetic maps. Genetic mapping for the traits segregating in the respective populations will facilitate the discovery of genomic regions controlling the phenotypic variation among peanut lines.
Acknowledgements
The authors would like to acknowledge Xuelin Luo for assisting with statistical analysis. The authors would like to acknowledge the funding support from the Georgia Peanut Commission, The Peanut Foundation, Feed the Future Innovation Lab for Collaborative Research on Peanut Productivity and Mycotoxin Control (Peanut and Mycotoxin Innovation Lab) supported by funding from the United States Agency for International Development (USAID), and the Agriculture and Food Research Initiative competitive grant 2012-85117-19435 of the USDA National Institute of Food and Agriculture.
Literature Cited
H.S., Atamian, P.A. Robertsand I. Kaloshian (2012). High and low throughput screens with root-knot nematodes Meloidogyne spp. J Vis Exp 61: 3629.
J.A. Baldwin, and J.P. Beasley (1990). Peanuts-A grower's guide to quality. Winston-Salem, NC: Planters LifeSavers Company. , .
R.W. Bauman, and A.J. Norden (1971). Effect of growth regulators on vegetative and reproductive characteristics of six peanut genotypes. J. Am. Peanut Res. Edu. Assoc 3: 75- 86.
Beasley, J.P. 2013 Planting dates In: University of Georgia 2013 Peanut Update Online at http://www.caes.uga.edu/commodities/fieldcrops/peanuts/documents/2013UGAPeanutProductionGuide.pdf
R. Bernardo, (2008). Molecular markers and selection for complex traits in plants: Learning from the last 20 years. Crop Sci 48: 1649- 1664.
D.J., Bertioli, S.B. Cannon, L. Froenicke, G. Huang, A.D. Farmer, E.K. Cannon, X. Liu, D. Gao, J. Clevenger, S. Dash, L. Ren, M.C. Moretzsohn, K. Shirasawa, W. Huang, B. Vidigal, B. Abernathy, Y. Chu, C. E. Niederhuth, P. Umale, A.C.G. Arajo, A. Kozik, K. D. Kim, M.D. Burrow, R.K. Varshney, X. Wang, X. Zhang, N. Barkley, P.M. Guimares, S. Isobe, B. Guo, B. Liao, H.T. Stalker, R.J. Schmitz, B.E. Scheffler, S.C.M. Leal-Bertioli, X. Xun, S.A. Jackson, R. Michelmore, and P. Ozias-Akins (2016). The genome sequences of Arachis duranensis and Arachis ipaensis, the diploid ancestors of cultivated peanut. Nat. Genet 48: 438- 446.
M.K., Beute, J.C. Wynneand D.A. Emery (1976). Registration of NC3033 peanut germplasm. Crop Sci 16: 887.
J.C., Biswas J.K. Ladha, F.B. Dazzo, Y.G. Yanni, B.G. Rolfe (2000). Rhizobial inoculation influences seedling vigor and yield of rice. Agron. J 92: 880- 886
W.D. Branch, and A.K. Culbreath (1995). Combination of early maturity and leaf spot tolerance within an advanced georgia peanut breeding line. Peanut Sci 22: 106- 108.
E. Buckler, and M. Gore (2007). An Arabidopsis haplotype map takes root. Nat. Genet 39: 1056- 1057.
E.S., Buckler, J.B. Holland, P.J. Bradbury, C.B. Acharya, P.J. Brown, C. Browne, E. Ersoz, S. Flint-Garcia, A. Garcia, J.C. Glaubitz, M.M. Goodman, C. Harjes, K. Guill, D.E. Kroon, S. Larsson, N.K. Lepak, H.H. Li, S.E. Mitchell, G. Pressoir, J.A. Peiffer, M.O. Rosas, T.R. Rocheford, M.C. Romay, S. Romero, S. Salvo, H.S. Villeda, H. S. da Silva, Q. Sun, F. Tian, N. Upadyayula, D. Ware, H. Yates, J.M. Yu, Z.W. Zhang, S. Kresovich, and M.D. McMullen (2009). The genetic architecture of maize flowering time. Science 325: 714- 718.
A.S. Bunting, (1955). A classification of cultivated groundnut. Empire J. Exp. Agri 23: 158- 170.
J., Clevenger, C. Chavarro, S.A. Pearl, P. Ozias-Akinsand S.A. Jackson (2015). Single nucleotide polymorphism identification in polyploids: A review, example, and recommendations. Mol. Plant 8: 831- 846.
J., Clevenger, Y. Chu, C. Chavarro, G. Agarwal, D.J. Bertioli, S.C.M. Leal-Bertioli, M.K. Pandey, J. Vaughn, B. Abernathy, N.A. Barkley, R. Hovav, M. Burrow, S.N. Nayak, A. Chitikineni, T. Isleib, C.C. Holbrook, S.A. Jackson, R.K. Varshney, and P. Ozias-Akins (2017). Genome-wide SNP genotyping resolves signatures of selection and tetrasomic recombination in peanut. Mol. Plant 10: 309- 322.
J.P. Clevenger, and P. Ozias-Akins (2015). SWEEP: A tool for filtering high-quality SNPs in polyploid crops. G3-Genes Genom Genet 5: 1797- 1803.
B.C.Y., Collard, M.Z.Z. Jahufer, J.B. Brouwerand E.C.K. Pang (2005). An introduction to markers, quantitative trait loci (QTL) mapping and marker-assisted selection for crop improvement: the basic concept. Euphytica 142: 169- 196.
M., Company, H.T. Stalkerand J.C. Wynne (1982). Cytology and leafspot resistance in Arachis hypogaea × wild species hybrids. Euphytica 31: 885- 893.
A.K. Culbreath, and R. Srinivasan (2011). Epidemiology of spotted wilt disease of peanut caused by tomato spotted wilt virus in the southeastern U.S. Virus Res 159: 101- 109.
P.M., Dang, C.Y. Chenand C.C. Holbrook (2012). Identification of genes encoding for drought-induced transcription factors in peanut (Arachis hypogaea L.). J. Mol. Biochem 1: 196- 205.
J.W., Dorner, R.J. Cole, T.H. Sandersand P.D. Blankenship (1989). Interrelationship of kernel water activity, soil-temperature, maturity, and phytoalexin production in preharvest aflatoxin contamination of drought-stressed peanuts. Mycopath 105: 117- 128.
I., Faye, M.K. Pandey, F. Hamidou, A. Rathore, O. Ndoye, V. Vadezand R.K. Varshney (2015). Identification of quantitative trait loci for yield and yield related traits in groundnut (Arachis hypogaea L.) under different water regimes in Niger and Senegal. Euphytica 206: 631- 647.
D.W. Gorbet, and B.L. Tillman (2009). Registration of 'Florida-07' peanut. J. Plant Reg 3: 14- 18.
B.H., Guo, D.A. Sleperand W.D. Beavis (2010). Nested association mapping for identification of functional markers. Genetics 186: 373- 383.
C.C. Holbrook, and A.K. Culbreath (2007). Registration of 'Tifrunner' Peanut. J. Plant Reg 1: 124.
C.C., Holbrook, T.G. Isleib, P. Ozias-Akins, Y. Chu, S. Knapp, B. Tillman, B. Guo., R. Gill, and M.D. Burow (2013). Development and phenotyping of recombinant inbred line (RIL) populations for peanut (Arachis hypogaea). Peanut Sci 40: 89- 94.
D.C.H. Hsi, and R.E. Finkner (1972). Registration of New Mexico Valencia A peanut. Crop Sci 12: 256.
T.G., Isleib, S.R. Milla-Lewis, H.E. Pattee, S.C. Copeland, M.C. Zuleta, B.B. Shew, J.E. Hollowell, T.H. Sanders, L.O. Dean, K.W. Hendrix, M. Balota, and J.W. Chapin (2011). Registration of 'Bailey' Peanut. J. Plant Reg 5: 27- 39.
S., Jain, S. Srivastava, N.B. Sarinand N.N.V. Kav (2006). Proteomics reveals elevated levels of PR 10 proteins in saline-tolerant peanut (Arachis hypogaea) calli. Plant Physiol. Bioch 44: 253- 259.
N., Jeong, J.K. Moon, H.S. Kim, C.G. Kimand S.C. Jeong (2011). Fine genetic mapping of the genomic region controlling leaflet shape and number of seeds per pod in the soybean. Theor. Appl. Genet 122: 865- 874.
D.L., Ketring, W.R. Jordan, D. Smithand C.E. Simpson (1982). Genetic variability in root and shoot growth characteristics of peanut. Peanut Sci 9: 68- 72.
Khera, P., M.K. Pandey, H. Wang, S.P. Feng, L.X. Qiao, A.K. Culbreath, S. Kale, J.P. Wang, C.C. Holbrook, W.J. Zhuang, R. K. and B.Z. Guo 2016 Mapping quantitative trait loci of resistance to tomato spotted wilt virus and leaf spots in a recombinant inbred line population of peanut (Arachis hypogaea L.) from SunOleic 97R and NC94022 PLoS One 11 doi:10.1371/journal.pone.0158452.
D.A., Knauft, D.W. Gorbetand A.J. Norden (1988). Yield and market quality of seven peanut genotypes as affected by leafspot disease and harvest date. Peanut Sci 15: 9- 13.
K.L., Kump, P.J. Bradbury, R.J. Wisser, E.S. Buckler, A.R. Belcher, M.A. Oropeza-Rosas, J.C. Zwonitzer, S. Kresovich, M.D. McMullen, D. Ware, P.J. Balint-Kurti, and J.B. Holland (2011). Genome-wide association study of quantitative resistance to southern leaf blight in the maize nested association mapping population. Nat. Genet 43: 163- U120.
O., Ladejobi, J. Elderfield, K.A. Gardner, R.C. Gaynor, J. Hickey, J.M. Hibberd, I.J. Mackay, and A.R. Bentley (2016). Maximizing the potential of multi-parental crop populations. Appl. Transl. Genom 11: 9- 17.
L. Lamari, ((2008)). ASSESS Imaging analysis software for plant disease quantification. MN USA: Am. Phytopath. Soc. St. Paul. , .
M.D., McMullen, S. Kresovich, H.S. Villeda, P. Bradbury, H.H. Li, Q. Sun, S. Flint-Garcia, J. Thornsberry, C. Acharya, C. Bottoms, P. Brown, C. Browne, M. Eller, K. Guill, C. Harjes, D. Kroon, N. Lepak, S.E. Mitchell, B. Peterson, G. Pressoir, S. Romero, M.O. Rosas, S. Salvo, H. Yates, M. Hanson, E. Jones, S. Smith, J.C. Glaubitz, M. Goodman, D. Ware, J.B. Holland, and E.S. Buckler (2009). Genetic properties of the maize nested association mapping population. Science 325: 737- 740.
I.L., Miller, A.J. Norden, D.A. Knauftand D.W. Gorbet (1990). Influence of maturity and fruit yield on susceptibility of peanut to cercosporidium personatum (late leafspot pathogen). Peanut Sci 17: 52- 58.
B., Ney, C. Duthionand E. Fontaine (1993). Timing of reproductive abortions in relation to cell-division, water-Content, and growth of pea-seeds. Crop Sci 33: 267- 270.
A.J., Norden, R.W. Lipscomband W.A. Carver (1969). Registration of Florunner peanut. Crop Sci 9: 850.
M.K., Pandey, A.W. Khan, V.K. Singh, M.K. Vishwakarma, Y. Shasidhar, V. Kumar, V. Garg, B.S. Bhat, A. Chitikineni, P. Janila, B. Guo, and R.K. Varshney (2016). QTL-seq approach identified genomic regions and diagonostic markers for rust and late leaf spot resistance in groundnut (Arachis hypogaea L.). Plant Biotech 15: 927- 941.
M.K., Pandey, E. Monyo, P. Ozias-Akins, X. Liang, P. Guimaraes, S.N. Nigam, H.D. Upadhyaya, P. Janila, X. Zhang, B. Guo, D.R. Cook, D.J. Bertioli, R. Michelmore, and R.K. Varshney (2012). Advances in Arachis genomics for peanut improvement. Biotech. Adv 30: 639- 651.
M.K., Pandey, H. Wang, P. Khera, M.K. Vishwakarma, S.M. Kale, A.K. Culbreath, C.C. Holbrook, X.J. Wang, R.K. Varshney, and B.Z. Guo (2017). Genetic dissection of novel QTLs for resistance to leaf spots and tomato spotted wilt virus in peanut (Arachis hypogaea L.). Front. Plant Sci 8: 25.
Rasheed, A., Y. Hao, X. Xia, A. Khan, Y. Xu, R.K. Varshney, and Z. He 2017 Crop breeding chips and genotyping platforms: progress, challenges and perspectives Mol. Plant doi:10.1016/j.molp.2017.06.008.
T.H. Sanders, (1980). Effects of variety and maturity on lipid class composition of peanut oil. J.Am. Oil Chem 57: 8- 11.
T.H., Sanders, J.R. Vercellotti, K.L. Crippenand G.V. Civille (1989). Effect of maturity on roast color and descriptive flavor of peanuts. J. Food Sci 54: 475- 477.
C.E., Simpson, M.R. Baring, A.M. Schubert, H.A. Melouk, Y. Lopezand J.S. Kirby (2003). Registration of 'OLin' peanut. Crop Sci 43: 1880- 1881.
J.Q., Shi, J.P. Zhan, Y.H. Yang, J. Ye, S.M. Huang, R.Y. Li, X.F. Wang, G.H. Liu, and H.Z. Wang (2015). Linkage and regional association analysis reveal two new tightly-linked major-QTLs for pod number and seed number per pod in rapeseed (Brassica napus L.). Sci. Rep 5: 14481.
J. Smartt, (1961). Groundnut varieties of Northern Rhodesia and their classification. Empire J. Exp. Agri 29: 153- 158.
V., Sujay, M.V. Gowda, M.K. Pandey, R.S. Bhat, Y.P. Khedikar, H.L. Nadaf, B. Gautami, C. Sarvamangala, S. Lingaraju, T. Radhakrishan, S.J. Knapp, and R.K. Varshney (2012). Quantitative trait locus analysis and construction of consensus genetic map for foliar disease resistance based on two recombinant inbred line populations in cultivated groundnut (Arachis hypogaea L.). Mol. Breed 30: 773- 788.
I.M. Sussex, and N.M. Kerk (2001). The evolution of plant architecture. Curr. Opin. Plant Biol 4: 33- 37.
S.P., Tallury, T.G. Isleib, S.C. Copeland, P. Rosas-Anderson, M. Balota, D. Singh, and H.T. Stalker (2014). Registration of two multiple disease-resistant peanut germplasm lines derived from Arachis cardenasii Krapov. & WC Gregory, GKP 10017. J.Plant Reg 8: 86- 89.
Tillman, B. L., Gorbet, D. W., Culbreath, A. K., and Todd, J. W. 2006 Response of peanut cultivars to seeding density and row patterns Crop Management doi:10.1094/CM-2006-0711-01-RS.
G.M., Timmerman-Vaughan, A. Mills, C. Whitfield, T. Frew, R. Butler, S. Murray, M. Lakeman, J. McCallum, A. Russell, Wilson (2005). Linkage mapping of QTL for seed yield, yield components, and developmental traits in pea. Crop Sci 45: 1336- 1344.
UGA (2018) Georgia Pest Management Handbook - Commercial Edition: College of Agricultural & Environmental Sciences Pp 848
R.K., Varshney, D.J. Bertioli, M.C. Moretzsohn, V. Vadez, L. Krishnamurthy, R. Aruna, S.N. Nigam, B.J. Moss, K. Seetha, K. Ravi, G. S.J. Knapp, and D.A. Hoisington (2009). The first SSR-based genetic linkage map for cultivated groundnut (Arachis hypogaea L.). Theor. Appl. Genet 118: 729- 739.
E.J. Williams, and J.S. Drexler (1981). A non-destructive method for determining peanut pod maturity. Peanut Sci 8: 134- 141.
E.J., Williams, G.O. Ware, J.-Y. Laiand J.S. Drexler (1987). Effect of pod maturity and plant age on pod and seed size distributions of Florrunner peanuts. Peanut Sci 14: 79- 83.
C., Wu, R. Gill, Y. Chu, C.C. Holbrookand P. Ozias-Akins (2015). Fine phenotyping of pod and seed traits in Arachis germplasm accessions using digital image analysis. Peanut Sci 42: 65- 73.
Zhou, X.J., Y.L. Xia, J.H. Liao, K.D. Liu, Q. Li, Y. Dong, X.P. Ren, Y.N. Chen, L. Huang, B.S. Liao, Y. Lei, L.Y.. Yan and H.F. Jiang 2016 Quantitative trait locus analysis of late leaf spot resistance and plant-type-related traits in cultivated peanut (Arachis hypogaea L.) under multi-environments PLoS One https://doi.org/10.1371/journal.pone.0166873
Notes
- Horticulture Department, University of Georgia Tifton Campus, Tifton, GA 31793; [^]
- USDA- Agricultural Research Service, Crop Genetics and Breeding Research Unit, Tifton, GA 31793; [^]
- Department of Crop Science, North Carolina State University, P.O. Box 7629, Raleigh, NC 27695; [^]
- Texas A&M AgriLife Research, Lubbock TX 79403, and Department of Plant and Soil Science, Texas Tech University, Lubbock, TX 79409; [^]
- Department of Plant Pathology, University of Georgia, Tifton, Georgia 31793; [^]
- North Florida Research and Extension Center, University of Florida, Marianna, FL 32446; [^]
- Institute of Plant Breeding, Genetics & Genomics, University of Georgia Tifton Campus, Tifton, GA 31793. [^] *Corresponding author's E-mail: pozias@uga.edu
- Current address: Shandong Peanut Research Institute, Qingdao, 266100 P.R. China [^]
- Current address: Mars Wrigley Confectionery, Center for Applied Genetic Technologies, Athens, GA, 30606 [^]
Author Affiliations