Introduction
Recent progress in sequencing Arachis genomes has highlighted the importance of developing or refining phenotyping methods for various traits including disease resistance (Brenneman et al., 2014; Chen et al., 2014; Culbreath et al., 2014). Genes contributing to disease resistance or other traits can only be verified by phenotyping peanut genotypes (Holbrook et al., 2014). Sclerotinia blight, caused by Sclerotinia minor Jagger, is one of the most important diseases of peanut worldwide and a priority for phenotyping. In the U.S., S. minor is of particular concern in the Virginia-Carolina and Southwest production areas where it may cause over 50% in yield losses (Porter and Melouk, 1997).
Field evaluations conducted over multiple years and in multiple locations are the ideal standard for evaluating resistance to diseases including Sclerotinia blight (Brenneman et al., 2014). Unfortunately, field trials are limited by time (season), space, variable soilborne pathogen distribution, and weather. Laboratory-based resistance assays can bypass many disadvantages of field tests, but laboratory assays also have unique limitations. Most significantly, laboratory resistance assays can only measure physiological resistance and do not assess the significant effect of plant canopy architecture on disease resistance (Brenneman et al., 1988; Bailey and Brune, 1997; Hollowell et al., 2003, 2008). However, markers associated with physiological resistance are generally the primary interest for phenotyping. In addition, fields may contain multiple genotypes of S. minor (Hollowell et al., 2003; Wu and Subbarao, 2006), but laboratory-based inoculation assays often use only one isolate of S. minor for logistical reasons, particularly if many peanut genotypes are being screened. Lastly, resistance assays may use plant parts, such as leaflets, considered less critical to the epidemiology of Sclerotinia blight. The relevance of these assays to field conditions may be a legitimate concern. Despite such limitations, laboratory-based assays may be valuable for phenotyping and for identifying susceptible germplasm for removal before commencing field trials.
Several methods for evaluating peanut genotypes in the laboratory for resistance to Sclerotinia blight have been published. Some assays used main stems and lateral branches of whole, intact plants (Goldman et al., 1995; Cruickshank et al., 2002; Woodward et al., 2006; Hollowell et al., 2008). Goldman et al. (1995) inoculated non-detached main stems of six-week-old plants with S. minor and found a significant relationship using a chi-square test between three years of field results and the laboratory assays. The assay was particularly useful in identifying susceptible entries: plants that were susceptible in the assay had an 87% chance of being susceptible in the field. However, plants identified as being resistant in the laboratory assay had only a 33% chance of being resistant in the field (Goldman et al., 1995). In Australia, Cruickshank et al. (2002) also reported a significant correlation between one year of field results and an assay similar to the one used by Goldman et al. In contrast, Hollowell et al. (2008) did not find that intact plant assays correlated well with four years of field data, but the correlation improved significantly when field data from a year with low disease pressure was removed from the analysis.
Several assays used excised main and lateral stems (Brenneman et al., 1988; Melouk et al., 1992; Chappell et al., 1995). Brenneman et al. (1988) inoculated various segments of detached lateral stems in moist chambers, and found that wounding was necessary for consistent infection. Mean lesion length was greater on stems inoculated at nodes where leaves had been removed than on wounds between nodes. Lesions were also longer on younger, terminal stem tissue than on older basal tissue. With the exception of one peanut genotype, Brenneman et al. obtained similar results using field-grown and greenhouse-grown plants (Brenneman et al., 1988). Melouk et al. (1992) inoculated detached shoot tips of main stems partially submersed in Hoagland’s solution. Significant differences were found in lesion length, and production and viability of sclerotia among 15 peanut genotypes. A complementary study found that the strength of the correlation between greenhouse and field resistance depended on the cultivar (Akem et al., 1992). Chappell et al. (1995) observed significant differences in lesion size among nine cultivars using detached lateral stems, but found more differences among cultivars using non-wounded than wounded stems. In contrast to Brenneman et al. (1988), cultivar differences in resistance were observed only on stems produced in the field, not the greenhouse. In addition, Chappell et al. did not find a correlation between results obtained from the detached stem assays to field resistance (Chappell et al., 1995).
Detached leaf assays have been used to identify quantitative trait loci for resistance to S. sclerotiorum in soybean (Arahana et al., 2001) and Brassica napus (Wu et al., 2013). In peanut, Hollowell et al. (2003) developed an excised leaflet assay after obtaining inconsistent results using detached stems. Leaflets from 8-week-old plants were placed on a screen mesh on top of moist sand and inoculated with 4-mm colonized agar plugs on the adaxial side. A moderate, but non-significant correlation between leaflet mean lesion lengths and field ratings was observed for twelve breeding lines (Hollowell et al., 2003). Smith et al. (2006) inoculated detached pegs without wounding in addition to detached leaflets, main stems, and vegetative and flowering nodes of lateral branches. Symptoms were more severe on pegs and leaves than on main stems or lateral branches, and cultivar by plant part interactions were observed. Woodward et al. (2006) used both detached leaflet and intact plants to evaluate eight peanut cultivars for resistance to S. sclerotiorum. Results from the two assays were highly correlated, but were not entirely consistent with field evaluations. The greenhouse assays identified the most susceptible and resistant cultivars, but for remaining cultivars, Woodward et al. (2006) hypothesized that variation in canopy structure and maturity contributed to the different results obtained from the field and greenhouse.
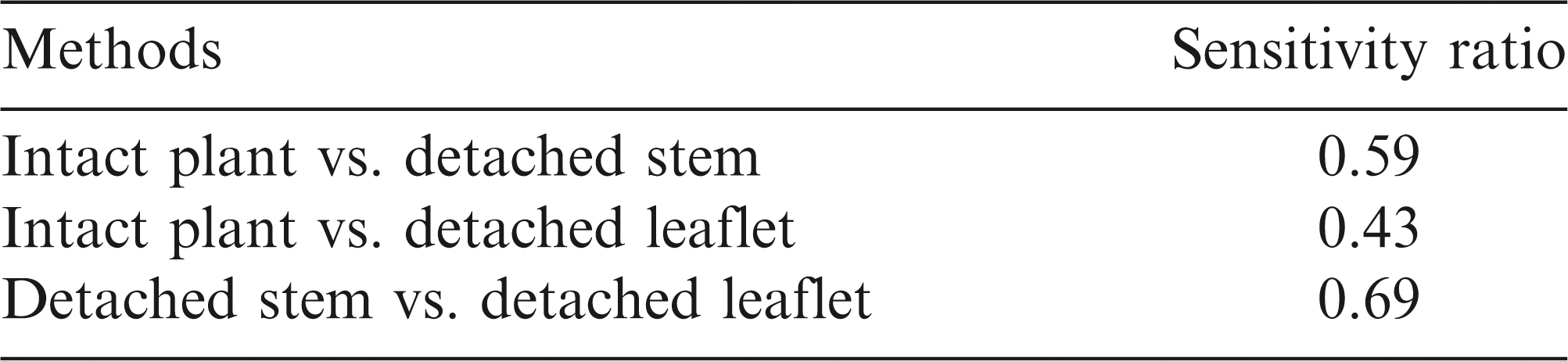
Determining the optimum resistance assay among methods which use different approaches can be problematic due to differences in scale and unknown relationships among these scales (Otto-Hanson et al., 2009). The sensitivity ratio, a statistical method that does not assume relationships between scales, can be used to identify the most efficient assay, i.e. the one with the least measurement variability (Kull et al., 2003; Otto-Hanson et al., 2009). The objectives of this project were to: (a) test three inoculation methods for evaluating resistance of six runner peanut genotypes to S. minor, and (b) determine the optimal inoculation method by comparing the three methods using the sensitivity ratio.
Materials and Methods
Six runner peanut genotypes were used to compare the three inoculation methods: the resistant cultivar Georgia-03L (Branch, 2004; Woodward et al., 2006) and breeding line ARSOK-R35 (Bennett and Chamberlin, 2014, 2015); moderately resistant cultivars Red River Runner (Melouk et al., 2012) and Tamrun 96 (Smith et al., 1998); and susceptible cultivars Okrun (Banks et al., 1989) and Tamrun OL02 (Simpson et al., 2006). Two seeds each were planted into a commercial potting soil (Metro-Mix 350, Sun Gro Horticulture, Agawam, MA) in 11 cm diam. (ca. 600 ml) pots for the intact plant and leaflet assays, or 14.8 cm diam. (ca. 1.67 L) pots for the detached plant assay. Pots were kept in a greenhouse maintained at 22-32 C under 300-watt LED lights (Grow Pro, LED Lighting for Less) set at 14 hr photophase. After emergence, the least vigorous seedling was removed by cutting the main stem below the soil line. The remaining seedling was grown for eight weeks and fertilized with 30 ml of 0.2% ammonium nitrate at five and seven weeks after planting. Six plants of each genotype were used in each experiment (inoculation method). One additional plant of each genotype was reserved for a non-inoculated control in each experiment.
One virulent isolate of S. minor (12-1), originally obtained from a diseased peanut stem in Fort Cobb, OK in 2012 was used. The isolate was maintained as sclerotia on colonized oat grains at 9 C (Chappell et al., 1995; Hollowell et al., 2003), and re-isolated annually from an inoculated peanut plant. Three to four days before plants were inoculated, 90 mm Petri dishes were filled with 15 ml of potato dextrose agar (PDA) using a peristaltic pump (UniSpense, Wheaton, Millville, NJ). Two-day-old cultures of S. minor were prepared by placing a single oat grain colonized with two or more sclerotia in the center of each Petri dish. Petri dishes were sealed with Parafilm (Bemis Co. Inc., Oshkosh, WI), and cultured in the dark in an incubator set at 20 C. Petri dishes for mock-inoculated controls were prepared the same way but were not inoculated with S. minor.
Intact Plant Assay
The six runner genotypes were arranged in randomized complete blocks on 35.6 cm × 45.7 cm cafeteria trays. Petri dishes with colony diameters of 4 cm or greater were used to inoculate plants, and one Petri dish was used to inoculate all plants within each block (tray). Plugs were cut from the actively growing margin of the colony with a sterile cork borer (5 mm diam.); plugs cut from sterile PDA were used for the mock-inoculated control plants. A sterile metal spatula was used to transfer each plug from the plate to a capless 0.2 ml PCR tube so that the mycelia-covered side of the plug was on the top of the tube. Desiccation of plugs was prevented by covering each tube with Parafilm and cutting a small incision (ca. 6 mm) in the Parafilm. The first true leaf that did not subtend a vegetative branch from the bottom of the main stem was removed with a razor blade so that approximately 15 cm of the petiole remained attached to main stem. Plants were inoculated by placing the tube over the severed petiole, making sure that the plug was in contact with the freshly cut surface.
Each block/tray was placed in a growth chamber (PGR15; Controlled Environments Ltd., Winnipeg, MB) set at 22 C. High humidity (98%) was maintained by lining the growth chamber shelf with saturated bath towels which were rewetted every three days. Additional humidity was provided by a Cyclone fog machine placed inside the chamber (FutureGarden, North Lindenhurst, NY). Plants received fluorescent light set at 14-hr photophase. The length of the lesion on the main stem was measured daily from three to seven days after inoculation using a digital caliper (Mitutoyo America, Aurora, IL). PCR tubes were removed from the plants if petioles collapsed after infection.
Detached Stem Assay
Detached main stems were prepared for inoculation by using a razor to remove the main stem immediately above the first node, cutting perpendicular to the stem. Side branches arising from the second node were also removed, using cuts parallel to the main stem. A sterile foam plug (1.8 cm long × 1.6 cm diam.), which was previously cut lengthwise ca. 1 cm deep, was used to secure the main stem into a 16 × 150 mm test tube (VWR, Radnor, PA) filled with 20 ml of sterile, ½-strength Hoagland’s nutrient solution (Sigma-Aldrich, St. Louis, MO; Barac et al., 2004). Approximately 2.5 cm of the stem was immersed in the solution.
A leaf from the next node not subtending a vegetative branch (usually the third node), was inoculated as described for the intact plant assay. Six test tubes containing one main stem of the six cultivars were arranged in a randomized complete block design in a 25 × 10 × 8 cm test tube rack. The six racks (blocks) were placed directly on the saturated bath towels in the growth chamber. A seventh rack contained stems mock-inoculated with PDA. Environmental conditions (temperature, light, humidity) were maintained, and lesion data were collected, as for the intact plant assay. Additional Hoagland’s solution was added to the test tubes as needed, generally two to three days after inoculation.
Detached Leaflet Assay
A modification of the detached leaflet assay of (Hollowell et al., 2003; Smith et al., 2006) was used. Two leaflets each from the second and third fully expanded leaves from the apex of the plant were used for the detached leaflet assay. The four leaflets were arranged in rows by cultivar on two layers of fiberglass screen mesh placed on top of approximately 7.6 cm of sterile moist sand in a container (42 × 29 × 14 cm) that had been wiped down with 70% reagent alcohol. The sand was prepared by autoclaving 22 kg of sand with 2 L of reverse-osmosis water in a covered container for 60 min. The container was autoclaved three consecutive days without adding water after the first day. The order of cultivar rows was randomly assigned for each of the six leaflet chambers/blocks. The center of each leaflet was inoculated using a sterile, 2 mm diam. amalgam carrier (Pulpdent, Watertown, MA; Porter, 2012) by placing a colonized PDA plug, taken from the colony margin, mycelium-side up on the middle of each leaflet. A seventh container held leaflets mock-inoculated with PDA.
Leaflet chamber boxes were placed in a growth chamber set at 22 C without light. Brown and necrotic lesions, rather than water-soaked areas, were measured daily on days 2 to 5 after inoculation. Lesions that were irregular in shape were measured by recording the longest length, following the protocol of Hollowell et al. (2003). Temperature and relative humidity were monitored in two of the seven leaflet chambers with HOBO U-10 data loggers (Onset, Bourne, MA). Leaflets with lesions covering the entire surface area were removed from the chamber boxes so that mycelium did not infect adjacent leaflets.
Experimental Design and Statistical Analyses
Experiments were conducted between 13 Dec. 2013 and 14 May 2014. One inoculation method was evaluated per week and the three methods were alternated sequentially. The intact plant assay and leaflet assays were conducted five times, and the detached stem assay was evaluated six times. Blocks were removed from the analyses if the plate used to inoculate each block contained a less aggressive culture, i.e. less than 50% of the leaflets or stems were infected.
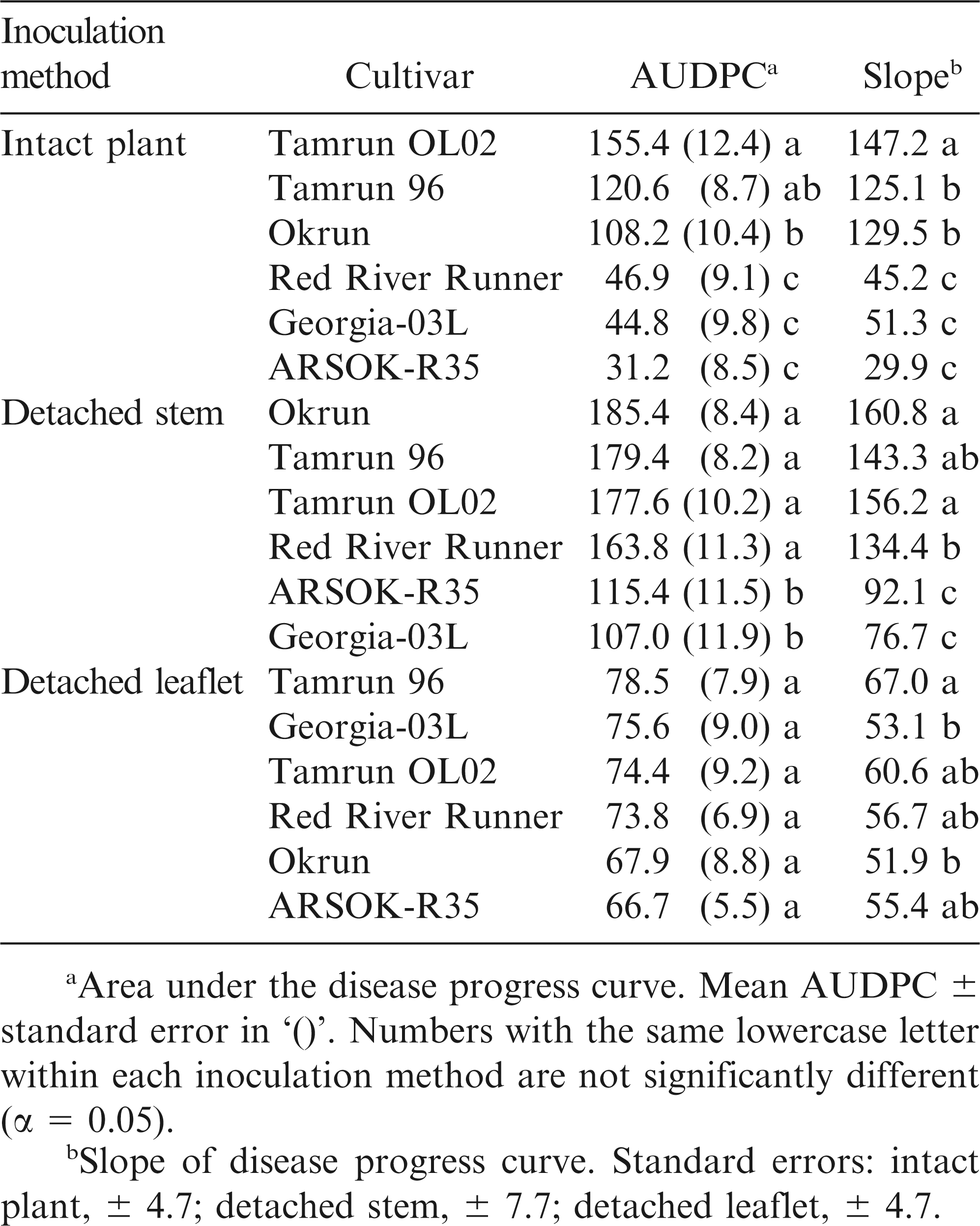
For the intact plant and detached stem assays, the response variable was stem lesion length. For the leaflet assay, mean lesion lengths from the four leaflets were analyzed. Each inoculation method was analyzed separately. Hypotheses regarding cultivar differences in resistance to Sclerotinia blight were tested using repeated measures ANOVA with an autoregressive covariance structure (TYPE = AR(1)) in PROC MIXED of SAS (SAS, ver. 9.3, SAS Institute, Cary, NC). When a relevant interaction was significant, the SLICE option of the LSMEANS statement was used to examine simple effects. In each set of multiple comparisons, experiment-wise type I error was controlled at α = 0.05 using ADJUST = TUKEY option. Disease progression for each cultivar within each inoculation method was estimated by calculating the area under the disease progress curve (AUDPC; Shaner and Finney, 1977) and compared using mixed-model ANOVA in PROC MIXED. Estimates of the slope of the length of lesion length over time were also estimated within ANOVA using orthogonal contrasts. The three inoculation methods were compared using the sensitivity ratio method of Otto-Hanson et al. (2009).
Results and Discussion
No symptoms were observed on the non-inoculated control plants for any of the inoculation methods. A total of three blocks each from the detached stem and detached leaflet assays were not included in the analyses due to poor infection rates. Although S. minor isolate 12-1 was cultured and maintained following published methods (Chappell et al., 1995; Hollowell et al., 2003), morphological changes and loss of pathogenicity in Sclerotinia cultures maintained in the laboratory were previously reported (Pratt, 2001).
Intact Plant Assay
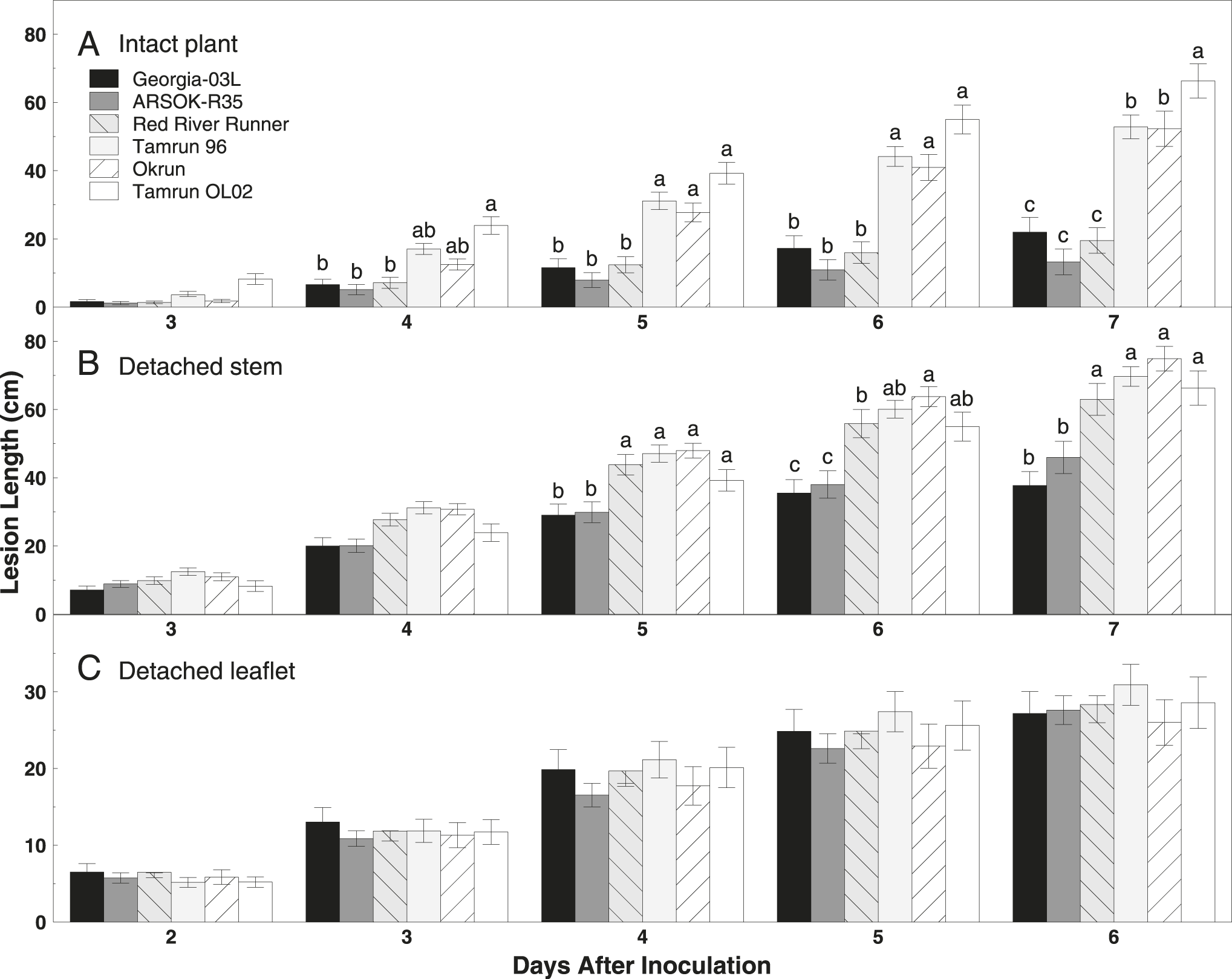
Lesion length was significantly affected by cultivar (F = 40.11; df = 5, 266; P < 0.01) and time (F = 320.79; df = 4, 754; P < 0.01), but a significant interaction between cultivar and time (F = 11.12; df = 20, 796; P < 0.01) indicated that the effect of one was dependent on the level of the other (Fig. 1). Smaller lesions were observed on main stems of Georgia-03L, ARSOK-R35, and Red River Runner than on Tamrun 96, Okrun, and Tamrun OL02 at five to seven days after inoculation (Fig. 1). By day seven, Tamrun OL02 had the longest lesions (Fig. 1).
Bar graph of mean lesion length (±SE) over time on runner peanut genotypes Georgia‐03L, ARSOK‐R35, Red River Runner, Tamrun 96, Okrun, and Tamrun OL02 inoculated with Sclerotinia minor using (A) intact plant, (B) detached stem, and (C) detached leaflet assays. Means followed by the same lowercase letter among cultivars within inoculation method and day are not significantly different (α=0.05).
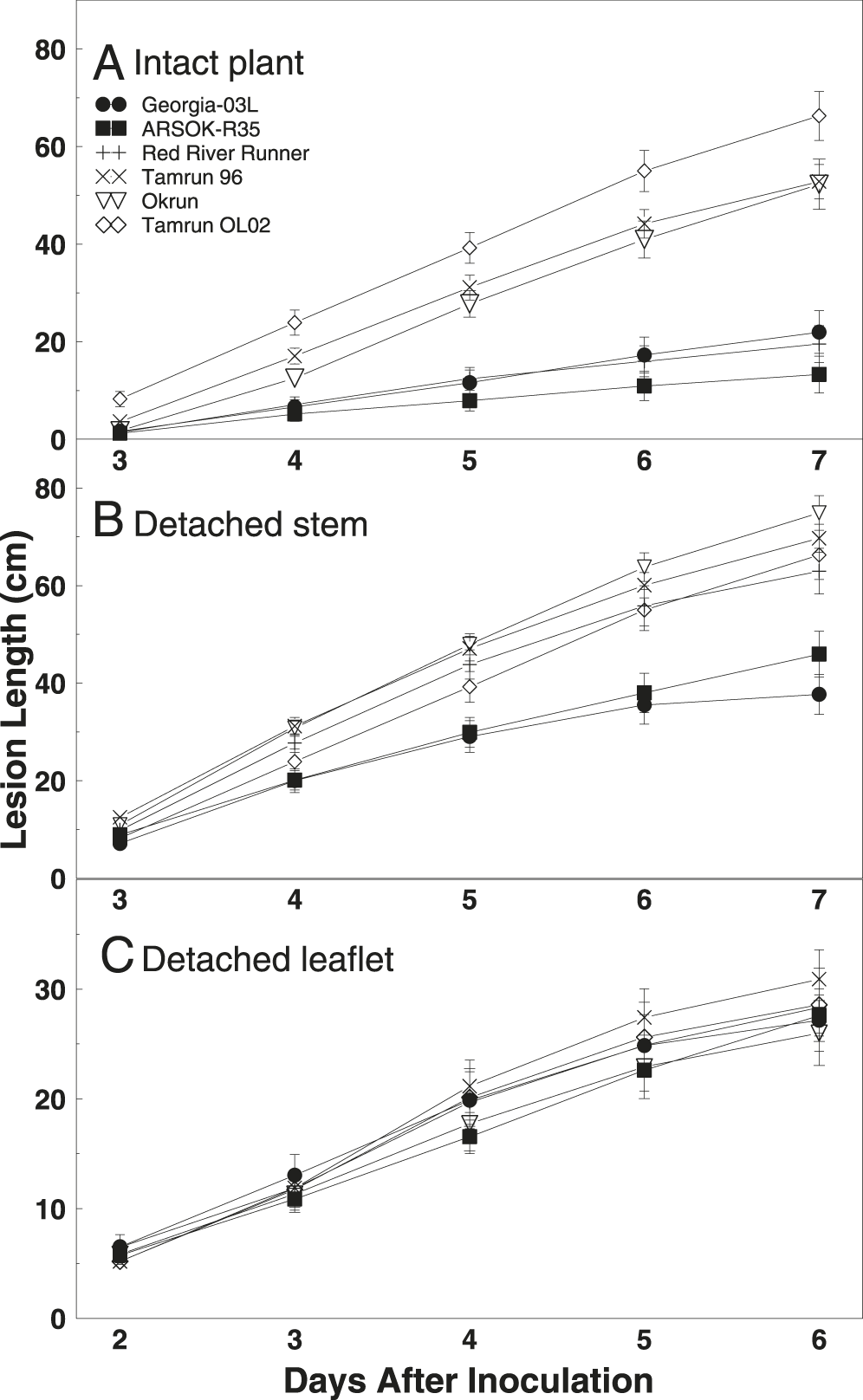
Differences in area under the disease progress curves were observed among cultivars (Fig. 2; F = 27.77; df = 5, 163; P < 0.01). AUDPC and rate of disease progress (slope) was smallest in ARSOK-R35, Georgia-03L, and Red River Runner (Table 1). Tamrun OL02 had the greatest AUDPC and rate of disease progress, and Okrun and Tamrun 96 were not significantly different from each other.
Detached Stem Assay
As observed in the intact plant assay, cultivar (F = 14.67; df = 5, 289; P < 0.01), time (F = 720.46; df = 4, 830; P < 0.01), and the interaction of cultivar and time (F = 6.81; df = 20, 880; P < 0.01) significantly affected lesion length in the detached stem assay. Differences in lesion length among cultivars were not observed until five days after inoculation (Fig. 1). On days five to seven, the most resistant cultivars were Georgia-03L and ARSOK-R35. In contrast to the results from the intact plant assay, in which Red River Runner was more similar to the most resistant cultivars, Red River Runner was among the susceptible cultivars in the detached stem assay (Figs. 1 and 2). Differences in AUDPC among cultivars were observed (Fig. 2; F = 13.76; df = 5, 182; P < 0.01). Rates of lesion growth for Georgia-03L and ARSOK-R35 were significantly less than for Red River Runner, Tamrun 96, Okrun, and Tamrun OL02 (Table 1). Okrun had the greatest rate of disease progression and AUDPC but did not differ significantly from Tamrun OL02 or Tamrun 96.
Detached Leaflet Assay
In the detached leaflet assay, lesion length was not affected by cultivar (Fig. 1; F = 0.45; df = 5, 259; P = 0.81), but increased over time (F = 378.50; df = 4, 698; P < 0.01). The interaction between the main effects of cultivar and time was not significant (F = 0.84; df = 4, 698; P = 0.66). No differences in AUDPC were observed among cultivars (Table 1; F = 0.46; df = 5, 147; P = 0.81), but rate of disease progress was numerically highest in Tamrun 96.
Sensitivity Ratio
Using the sensitivity ratio, the intact plant assay was the most efficient assay of the three inoculation methods. Approximately three ( = [1/0.59]2; Otto-Hanson et al., 2009) times as many replications are needed for the detached stem assay to equal the sensitivity of the intact plant assay (Table 2). The detached leaflet assay was the least sensitive method, requiring five ( = [1/0.43]2) times as many replications to equal the intact plant and twice ( = [1/0.69]2) as many replications as the detached stem assay.
In this study, the detached leaflet assay was unable to discriminate among the peanut entries even though lesions on all cultivars increased in size over time. Others have found the detached leaflet assay useful for identifying resistant genotypes (Hollowell et al., 2003; Smith et al., 2006; Woodward et al., 2006). One possible explanation for this discrepancy is the few peanut genotypes in common between this study and other studies.
Results from the intact plant and detached stem assays were relatively similar and both reflected resistance levels observed in the field better than the detached leaflet assay. Georgia-03L and ARSOK-R35, peanut entries known to be resistant in the field, had the smallest lesions and slowest disease progression. However, Red River Runner, a cultivar that is intermediate in resistance to Sclerotinia blight in the field, grouped among the resistant cultivars in the intact plant assay, but was more similar to susceptible entries in the detached stem assay. In another pathosystem, different responses were observed in assays using detached plant parts than in assays using intact plants. The hemibiotrophic pathogens Colletotrichum linicola and C. higginsianum caused more severe symptoms on detached than on attached leaves of Arabidopsis (Liu et al., 2007). The mechanisms responsible for the varying results with Red River Runner in this study using S. minor, a necrotroph, are unknown. Both the intact plant and detached stem assays used plants of the same age and inoculated the same plant part (petiole on main stem).
When pairwise comparisons were made among the three methods using the sensitivity ratio, the intact plant assay had the smallest error variance and required the fewest replications than the detached stem and leaflet assays. Although precise times for setup and maintenance were not recorded, the intact plant assay also requires less time than the detached stem assay because test tubes of Hoagland’s solution do not need to be prepared or refilled. Although the intact plant assay requires more space and is destructive, this method may be preferable if seed availability is not strictly limited.
Acknowledgements
The authors thank Angela Harting for technical assistance. This research was supported by USDA-ARS CRIS Project No. 3072-21220-007-00D. Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture. USDA is an equal opportunity provider and employer.
Literature Cited
Akem C.N. Melouk H.A. and Smith O.D. 1992 Field evaluation of peanut genotypes for resistance to Sclerotinia blight Crop Prot. 11 : 345 – 348 .
Arahana V.S. Graef G.L. Specht J.E. Steadman J.R. and Eskridge K.M. 2001 Identification of QTLs for resistance to in soybean Crop Sci. 41 : 180 – 188 .
Bailey J.E., and Brune P.D. 1997 Effect of crop pruning on Sclerotinia blight of peanut Plant Dis. 81 : 990 – 995 .
Banks D.J. Kirby J.S. and Sholar J.R. 1989 Registration of “Okrun” peanut Crop Sci. 29 : 1574 – 1574 .
Barac T. Taghavi S. Borremans B. Provoost A. Oeyen L. Colpaert J.V. Vangronsveld J. and van der Lelie D. 2004 Engineered endophytic bacteria improve phytoremediation of water-soluble, volatile, organic pollutants Nat. Biotechnol. 22 : 583 – 588 .
Bennett R.S. and Chamberlin K.D. 2014 Disease evaluations and agronomic traits of advance peanut breeding lines in 2013 . In Partners in Progress – Peanut Research at Oklahoma State University Oklahoma Agricultural Experiment Station, P-1044 , 13 – 16 .
Bennett R.S. and Chamberlin K.D. 2015 Disease evaluations and agronomic traits of advanced peanut breeding lines in 2014 . In Partners in Progress – Peanut Research at Oklahoma State University Oklahoma Agricultural Experiment Station, P-1042, 11 – 16 .
Branch W.D. 2004 Registration of “Georgia-03L” peanut Crop Sci. 44 : 1485 – 1486 .
Brenneman T.B. Phipps P.M. and Stipes R.J. 1988 A rapid method for evaluating genotype resistance, fungicide activity, and isolate pathogenicity of Sclerotinia minor in peanut Peanut Sci. 15 : 104 – 107 .
Brenneman T.B. Tillman B.L. and Dufault N. 2014 Phenotyping peanut diseases caused by soilborne pathogens Proc. Amer. Peanut Res. Educ. Soc. 46 : 16 – 17 (abstr .)
Chappell G.F. Shew B.B. Ferguson J.M. and Beute M.K. 1995 Mechanisms of resistance to Sclerotinia minor in selected peanut genotypes Crop Sci. 35 : 692 – 696 .
Chen C.Y. Butts C.L. Dang P.M. and Wang M.L. 2014 Advances in phenotyping of functional traits in the field crops Proc. Amer. Peanut Res. Educ. Soc. 46 : 15 (abstr .)
Cruickshank A.W. Cooper M. and Ryley M.J. 2002 Peanut resistance to Sclerotinia minor and S. sclerotiorum Aust. J. Agric. Res. 53 : 1105 – 1110 .
Culbreath A.K. Holbrook C.C. Guo B. Ozias-Akins P. Chu Y. Gilli R. Clevenger J. Brenneman T.B. and Isleib T.G. 2014 Phenotyping for foliar disease resistance Proc. Amer. Peanut Res. Educ. Soc. 46 : 16 (abstr .)
Goldman J.J. Smith O.D. Simpson C.E. and Melouk H.A. 1995 Progress in breeding Sclerotinia blight-resistant runner-type peanut Peanut Sci. 22 : 109 – 113 .
Holbrook C.C. Isleib T.G. Ozias-Akins P. Chu Y. Knapp S.J. Tillman B. Guo B. Barkley N. Chen C. and Burow M.D. 2014 Genetic resources for phenotyping Proc. Amer. Peanut Res. Educ. Soc. 46 : 15 – 16 (abstr .)
Hollowell J.E. Isleib T.G. Tallury S.P. Copeland S.C. and Shew B.B. 2008 Screening of Virginia-type peanut breeding lines for resistance to Cylindrocladium black rot and Sclerotinia blight in the greenhouse Peanut Sci. 35 : 18 – 24 .
Hollowell J.E. Shew B.B. and Isleib T.G. 2003 Evaluating isolate aggressiveness and host resistance from peanut leaflet inoculations with Sclerotinia minor Plant Dis. 87 : 402 – 406 .
Kull L.S. Vuong T.D. Powers K.S. Eskridge K.M. Steadman J.R. and Hartman G.L. 2003 Evaluation of resistance screening methods for Sclerotinia stem rot of soybean and dry bean Plant Dis. 87 : 1471 – 1476 .
Liu G. Kennedy R. Greenshields D.L. Peng G. Forseille L. Selvaraj G. and Wei Y. 2007 Detached and attached Arabidopsis leaf assays reveal distinctive defense responses against hemibiotrophic Colletotrichum spp Mol. Plant. Microbe Interact. 20 : 1308 – 1319 .
Melouk H.A. Akem C.N. and Bowen C. 1992 A detached shoot technique to evaluate the reaction of peanut genotypes to Sclerotinia minor Peanut Sci. 19 : 58 – 62 .
Melouk H.A. Chamberlin K. Godsey C.B. Damicone J. Burow M.D. Baring M.R. Simpson C.E. Dashiell K.E. and Payton M. 2012 Registration of “Red River Runner” Peanut J. Plant Regist. 7 : 22 – 25 .
Otto-Hanson L. Eskridge K.M. Steadman J.R. and Madisa G. 2009 The sensitivity ratio: a superior method to compare plant and pathogen screening tests Crop Sci. 49 : 153 – 160 .
Porter L. 2012 Selection of pea genotypes with partial resistance to Sclerotinia sclerotiorum across a wide range of temperatures and periods of high relative humidity Euphytica 186 : 1 – 8 .
Porter D.M. and Melouk H.A. 1997 Sclerotinia blight . p. 34 – 36 In: Kokalis-Burelle N. Porter D.M. Rodríguez-Kábana R. Smith D.H. and Subrahmanyam P. (eds .) Compendium of Peanut Diseases 2nd edition APS Press , St. Paul, MN .
Pratt R.G. 2001 Sclerotinia p. 74 – 78 In: Singleton L.L. Mihail J.D. & Rush C.M. (eds .) Methods for Research on Soilborne Phytopathogenic Fungi APS Press , St. Paul, MN .
Shaner G. and Finney R.E. 1977 The effect of nitrogen fertilization on the expression of slow-mildewing resistance in Knox wheat Phytopathology 67 : 1051 – 1056 .
Simpson C.E. Baring M.R. Schubert A.M. Black M.C. Melouk H.A. and Lopez Y. 2006 Registration of “Tamrun OL 02” peanut Crop Sci. 46 : 1813 .
Smith D.L. Hollowell J.E. Isleib T.G. and Shew B.B. 2006 Analysis of factors that influence the epidemiology of Sclerotinia minor on peanut Plant Dis. 90 : 1425 – 1432 .
Smith O.D. Simpson C.E. Black M.C. and Besler B.A. 1998 Registration of “Tamrun 96” peanut Crop Sci. 38 : 1403 .
Woodward J.E. Brenneman T.B. Kemerait Jr., A.K. Culbreath R.C. and Clark J.R. 2006 Sclerotinia blight in Georgia and evidence for resistance to Sclerotinia sclerotiorum in runner peanuts Plant Health Prog. Available at http://www.plantmanagementnetwork.org/pub/php/research/2006/runner/ .
Wu J. Cai G. Tu J. Li L. Liu S. Luo X. Zhou L. Fan C. and Zhou Y. 2013 Identification of QTLs for resistance to Sclerotinia stem rot and BnaC.IGMT5.a as a candidate gene of the major resistant QTL SRC6 in Brassica napus PLoS ONE 8 : e67740 .
Wu B.M. and Subbarao K.V. 2006 Analyses of lettuce drop incidence and population structure of Sclerotinia sclerotiorum and S. minor Phytopathology 96 : 1322 – 1329 .
Notes
- First and third authors: Research Plant Pathologist and Research Biologist, USDA-ARS, Wheat, Peanuts and Other Field Crops Research Unit, Stillwater, OK 74075. Second author: Professor, Department of Statistics, Oklahoma State University, Stillwater, OK 74078. *Corresponding author’s E-mail: rebecca.bennett@ars.usda.gov
Author Affiliations