Introduction
Root-knot nematodes (RKN) [Meloidogyne arenaria (Neal) Chitwood race 1] are a costly peanut (Arachis hypogaea L.) production problem in the U.S. (Kokalis-Burelle and Rodriguez-Kabana 1997; Starr et al., 2002). Introgression of RKN resistance from the wild species resulted in the first improved peanut cultivar ‘COAN’ (Simpson and Starr 2001) with a very high RKN resistance. Since then, several other RKN-resistant cultivars have been developed from COAN:‘NemaTAM’ (Simpson et al. 2003), ‘Webb’ (Simpson et al. 2013); ‘Tifguard’ (Holbrook et al. 2008); ‘Georgia-14N’ (Branch and Brenneman 2015), and ‘TifNV-High O/L’.
Inheritance of the very high RKN resistance found in COAN was first reported to be controlled as a single dominant gene in several studies (Burow et al. 1996; Choi et al. 1999; Chu et al. 2011; Church et al. 2000). Nagy et al. (2010) proposed the name Rma for this single dominant RKN resistance gene.
In 1996, Garcia et al. suggested two dominant genes (Mae and Mag) controlling egg number and galling, respectively. However, the F2 data were derived from different parental material than COAN. In yet another genetic study by Church et al. (2005), a recessive gene in addition to the single dominant gene was also proposed involving TxAG-6 (Simpson et al., 1993), the interspecific hybrid that served as the source of RKN resistance. The objective of current genetic study was to further determine the inheritance of a proposed second RKN resistance gene found in peanut derived from COAN.
Materials and Methods
During 2011 and 2012, a new field site at the Gibbs Farm (latitude: 31.4312°N and longitude: 83.5850°W) near the Coastal Plain Experiment Station, Tifton, GA was artificially inoculated with M. arenaria race 1. The susceptible peanut cultivar, ‘Georgia-10T’ (Branch and Culbreath 2011) was used to uniformly increase the peanut-specific race 1 nematode population during the growing season; whereas, hairy common vetch (Vicia villosa Roth) was used for the same purpose each winter as a susceptible cover crop.
Crosses were made in the greenhouse between the RKN-susceptible parent ‘Georgia Greener’ (Branch 2007) and two advanced RKN-resistant Georgia breeding lines, GA 082524 and GA 082546 (Branch et al. 2014). These two homozygous breeding lines resulted from the same three-way cross combination ‘Georgia-02C’ (Branch 2003) × [‘Georgia-01R’ (Branch 2002) × COAN].
Seed of the F2 and F3 cross populations were space-planted 30.5-cm apart during 2013 and 2014, respectively in the newly RKN inoculated field site. Planting dates were 22 May 2013 and 23 May 2014. Recommended cultural practices with irrigation were used throughout each growing season, except that no nematicides were applied with activity against RKN. The severity of nematode galling on roots and pods was rated after digging and inverting the plants each year. All plants were dug at the same time on 21 Oct 2013 and 29 Oct 2014. The percent damage was visually estimated from 0 to 100%, with 0% representing no galls and 100% representing galling on all pods and roots. Since there can sometimes be a low level of gall formation on resistant plants, a 5% level of galling was used as a threshold in defining resistant and susceptible genotypes. Data from visual classification of segregating RKN-resistant and RKN-susceptible plants and progeny row were analyzed by a chi-square program to test goodness-of-fit of observed vs expected genetic ratios.
Results and Discussion
During 2013, the F2 plant segregation showed an acceptable fit to both a 3:1 and 13:3, resistant to susceptible genetic ratios, respectively (Table 1). However, the 13:3 ratio had the best fit with the lowest chi-square values and highest probability compared to the 3:1 ratio.
Because of the number of genes inherited, one of the major differences between a 3:1 vs 13:3 ratio is that the F2 susceptible plants should not segregate for resistance and susceptible plants from a 3:1 ratio in the following F3 generation; whereas, the F2 susceptible plants would segregate from the 13:3 ratio. Both resistant and susceptible plants were found segregating within some of the F2:3 susceptible progeny rows in a 1 resistant to 3 susceptible ratio.
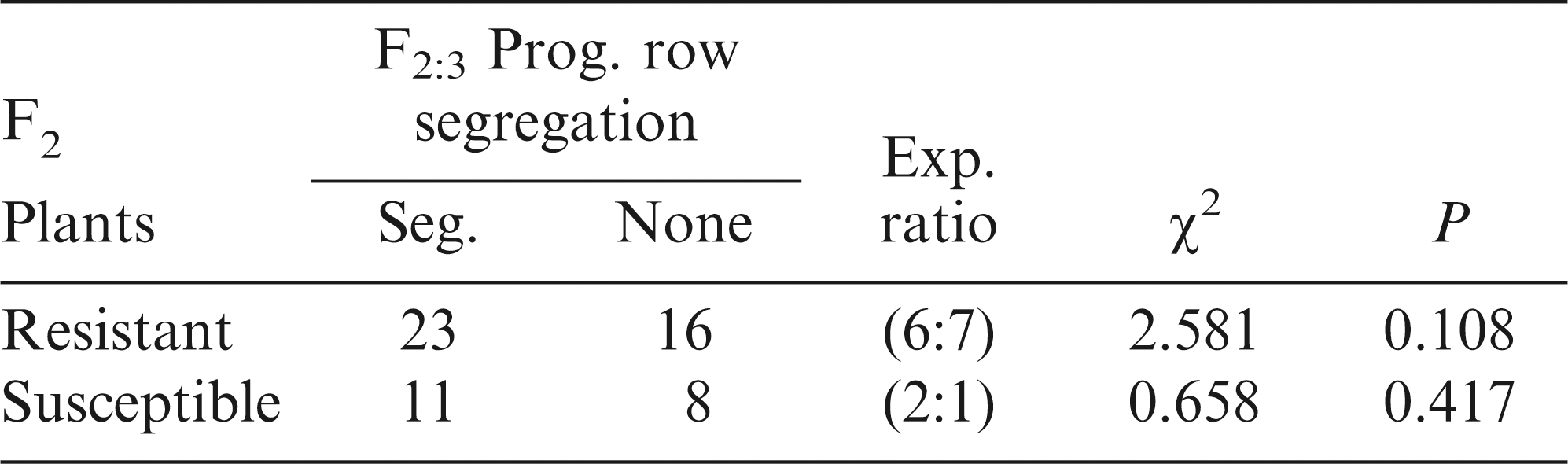
The F3 progeny row segregation showed an acceptable fit to a 6 segregating to 7 non-segregating ratio for the F2:3 RKN-resistant plants from the Georgia Greener x GA 082524 cross combination (Table 2). Likewise, the F3 progeny row segregation from the F2:3 RKN-susceptible plants was found to have an acceptable fit to a 2 segregating to 1 non-segregating ratio as expected for a 13:3 genetic model.
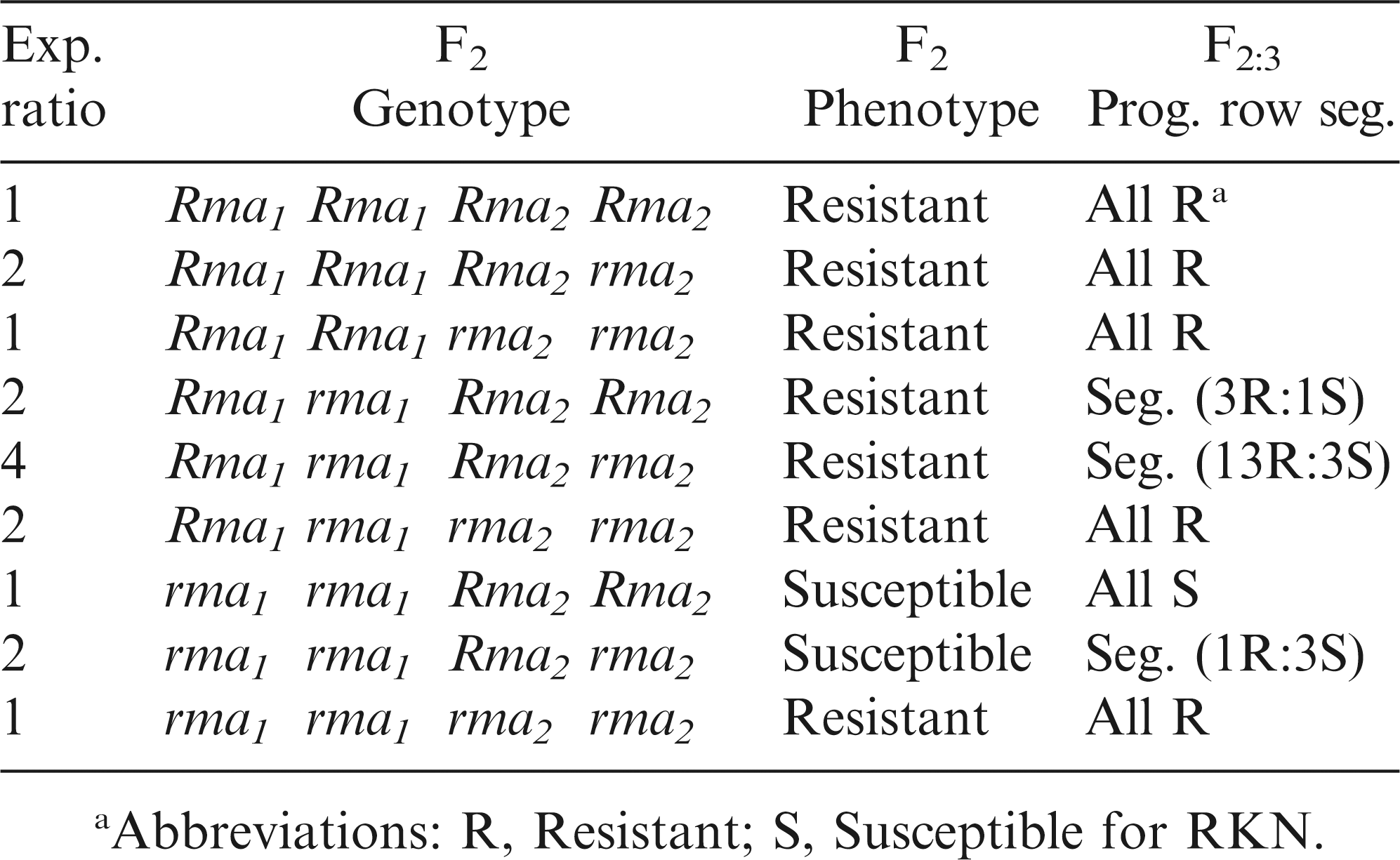
The evidence from this study strongly confirms the report by Church et al. (2005). The first RKN-resistant gene should be dominant as previously reported, designated Rma1 . However, the second RKN-resistance gene should be recessive with a proposed gene symbol, rma2 . As expected among segregating F2 plants and F2:3 progeny rows in a 13:3 ratio from a two-gene model (Table 3), either the homozygous or heterozygous dominant gene Rma1 Rma1 or Rma1 rma1 , homozygous recessive gene rma2 rma2 , or the various combinations of these two genes results in the very high level of peanut RKN resistance.
Literature Cited
Branch W.D. 2002 Registration of ‘Georgia-01R’ peanut Crop Sci. 42 : 1750 - 1751 .
Branch W.D. 2003 Registration of ‘Georgia-02C’ peanut Crop Sci. 43 : 1883 - 1884 .
Branch W.D. 2007 Registration of ‘Georgia Greener’ peanut J. Plant Reg. 1 : 121 .
Branch W.D. and Brenneman T.B. 2015 Registration of ‘Georgia-14N’ peanut J. Plant Reg. 9 : 159 - 161 .
Branch W.D. and Culbreath A.K. 2011 Registration of ‘Georgia-10T’ peanut J. Plant Reg. 5 : 279 - 281 .
Branch W.D. Brenneman T.B. and Hookstra G. 2014 Field test results versus marker assisted selection for root-knot nematode resistance in peanut Peanut Sci. 41 : 85 - 89 .
Burow M.D. Simpson C.E. Paterson A.H. and Starr J.L. 1996 Identification of peanut (Arachis hypogaea L.) RAPD markers diagnostic of root-knot nematode (Meloidogyne arenaria (Neal) Chitwood) resistance Mol. Breeding 2 : 369 - 379 .
Choi K. Burow M.D. Church G. Burow G. Paterson A.H. Simpson C.E. and Starr J.L. 1999 Genetics and mechanism of resistance to Meloidogyne arenaria in peanut germplasm J. Nematology 31 : 283 - 290 .
Church G.T. Simpson C.E. Burow M.D. Paterson A.H. and Starr J.L. 2000 Use of RFLP markers for identification of individuals homozygous for resistance to Meloidogyne arenaria in peanut Nematology 2 : 575 - 580 .
Church G.T. Starr J.L. and Simpson C.E. 2005 A recessive gene for resistance to Meloidogyne arenaria in interspecific Arachis spp hybrids. J. Nematology 37 : 178 - 184 .
Chu Y. Wu C.L. Holbrook C.C. Tillman B.L. Person G. and Ozias-Akins P. 2011 Marker-assisted selection to pyramid nematode resistance and the high oleic trait in peanut The Plant Genome. 4 : 110 - 117 .
Garcia G.M. Stalker H.T. Shroeder E. and Kochert G. 1996 Identification of RAPD, SCAR, and RFLP markers tightly linked to nematode resistance genes introgressed from Arachis cardenasii into Arachis hypogaea Genome 39 : 836 - 845 .
Holbrook C.C. Timper P. Culbreath A.K. and Kvien C.K. 2008 Registration of ‘Tifguard’ peanut J. Plant Reg. 2 : 92 - 94 .
Kokalis-Burelle N. and Rodriguez-Kabana R. 1997 Root-knot nematodes In: Compendium of Plant Diseases 2nd Ed., Amer. Phytopath. Soc. Press, pp. 45-48 .
Nagy E.D. Chu Y. Guo Y. Khanal S. Tang S. Li Y. Dong W.B. Timper P. Taylor C. Ozias-Akins P. Holbrook C.C. Beilinson V. Nielson N.C. Stalker H.T. and Knapp S.J. 2010 Recombination is suppressed in an alien introgression in peanut harboring Rma, a dominant root-knot nematode resistance gene Mol. Breeding 26 : 357 - 370 .
Simpson C.E. and Starr J.L. 2001 Registration of ‘COAN’ peanut Crop Sci. 41 : 918 .
Simpson C.E. Nelson S.C. Starr J.L. Woodard K.E. and Smith O.D. 1993 Registration of Tx-AG-6 and TxAG-7 peanut germplasm lines Crop Sci. 33 : 1418 .
Simpson C.E. Starr J. L Church G.T. Burow M.D. and Paterson A.H. 2003 Registration of ‘NemaTAM’ peanut Crop Sci. 43 : 1561 .
Simpson C.E. Starr J. L Baring M.R. Burow M.D. Cason J.M. and Wilson J.N. 2013 Registration of ‘Webb’ peanut J. Plant Reg. 7 : 265 - 268 .
Starr J.L. Bridge J. and Cook R. 2002 Resistance to plant-parasitic nematodes: history, current use and future potential . Pp: 1 - 22 In: Starr J. L. Cook R. and Bridge J. (eds). Plant Resistance to Parasitic Nematodes CABI Publishing, CAB International , New York, NY .
Notes
- Professor, University of Georgia, Dept. of Crop & Soil Sciences, Coastal Plain Experiment Station, 2360 Rainwater Road, Tifton, GA 31793-5766
- Professor, University of Georgia, Dept. of Plant Pathology, Coastal Plain Experiment Station, 2360 Rainwater Road, Tifton, GA 31793-5766
- Professor, University of Georgia, Dept. of Plant Pathology, Athens, GA 30602 *Corresponding author email: wdbranch@uga.edu
Author Affiliations